Complex rearranged small supernumerary marker chromosomes (sSMC), three new cases; evidence for an underestimated entity?
- PMID: 18471318
- PMCID: PMC2375881
- DOI: 10.1186/1755-8166-1-6
Complex rearranged small supernumerary marker chromosomes (sSMC), three new cases; evidence for an underestimated entity?
Abstract
Background: Small supernumerary marker chromosomes (sSMC) are present ~2.6 x 106 human worldwide. sSMC are a heterogeneous group of derivative chromosomes concerning their clinical consequences as well as their chromosomal origin and shape. Besides the sSMC present in Emanuel syndrome, i.e. der(22)t(11;22)(q23;q11), only few so-called complex sSMC are reported.
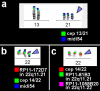
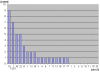
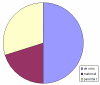
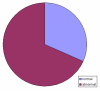
Results: Here we report three new cases of unique complex sSMC. One was a de novo case with a dic(13 or 21;22) and two were maternally derived: a der(18)t(8;18) and a der(13 or 21)t(13 or 21;18). Thus, in summary, now 22 cases of unique complex sSMC are available in the literature. However, this special kind of sSMC might be under-diagnosed among sSMC-carriers.
Conclusion: More comprehensive characterization of sSMC and approaches like reverse fluorescence in situ hybridization (FISH) or array based comparative genomic hybridization (array-CGH) might identify them to be more frequent than only ~0.9% among all sSMC.
Figures


References
-
- Liehr T, Mrasek K, Weise A, Dufke A, Rodriguez L, Martinez Guardia N, Sanchis A, Vermeesch JR, Ramel C, Polityko A, Haas OA, Anderson J, Claussen U, von Eggeling F, Starke H. Small supernumerary marker chromosomes–progress towards a genotype-phenotype correlation. Cytogenet Genome Res. 2006;112:23–34. doi: 10.1159/000087510. - DOI - PubMed
-
- Liehr T, Weise A. Frequency of small supernumerary marker chromosomes in prenatal, newborn, developmentally retarded and infertility diagnostics. Int J Mol Med. 2007;19:719–731. - PubMed
LinkOut - more resources
Full Text Sources

