Endogenous siRNA and miRNA targets identified by sequencing of the Arabidopsis degradome
- PMID: 18472421
- PMCID: PMC2583427
- DOI: 10.1016/j.cub.2008.04.042
Endogenous siRNA and miRNA targets identified by sequencing of the Arabidopsis degradome
Abstract
MicroRNAs (miRNAs) regulate the expression of target mRNAs in plants and animals [1]. Plant miRNA targets have been predicted on the basis of their extensive and often conserved complementarity to the miRNAs [2-4], as well as on miRNA overexpression experiments [5]; many of these target predictions have been confirmed by isolation of the products of miRNA-directed cleavage. Here, we present a transcriptome-wide experimental method, called "degradome sequencing," to directly detect cleaved miRNA targets without relying on predictions or overexpression. The 5' ends of polyadenylated, uncapped mRNAs from Arabidopsis were directly sampled, resulting in an empirical snapshot of the degradome. miRNA-mediated-cleavage products were easily discerned from an extensive background of degraded mRNAs, which collectively covered the majority of the annotated transcriptome. Many previously known Arabidopsis miRNA targets were confirmed, and several novel targets were also discovered. Quantification of cleavage fragments revealed that those derived from TAS transcripts, which are unusual in their production of abundant secondary small interfering RNAs (siRNAs), accumulated to very high levels. A subset of secondary siRNAs are also known to direct cleavage of targets in trans[6]; degradome sequencing revealed many cleaved targets of these trans-acting siRNAs (ta-siRNAs). This empirical method is broadly applicable to the discovery and quantification of cleaved targets of small RNAs without a priori predictions.
Figures



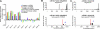
References
-
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. - PubMed
-
- Allen E, Xie Z, Gustafson AM, Carrington JC. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell. 2005;121:207–221. - PubMed
-
- Jones-Rhoades MW, Bartel DP. Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol. Cell. 2004;14:787–799. - PubMed
-
- Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B, Bartel DP. Prediction of plant microRNA targets. Cell. 2002;110:513–520. - PubMed
-
- Schwab R, Palatnik JF, Riester M, Schommer C, Schmid M, Weigel D. Specific effects of microRNAs on the plant transcriptome. Dev. Cell. 2005;8:517–527. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

