The tight junction protein complex undergoes rapid and continuous molecular remodeling at steady state
- PMID: 18474622
- PMCID: PMC2386107
- DOI: 10.1083/jcb.200711165
The tight junction protein complex undergoes rapid and continuous molecular remodeling at steady state
Abstract
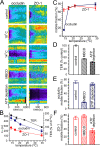
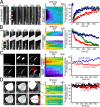
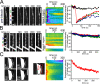
The tight junction defines epithelial organization. Structurally, the tight junction is comprised of transmembrane and membrane-associated proteins that are thought to assemble into stable complexes to determine function. In this study, we measure tight junction protein dynamics in live confluent Madin-Darby canine kidney monolayers using fluorescence recovery after photobleaching and related methods. Mathematical modeling shows that the majority of claudin-1 (76 +/- 5%) is stably localized at the tight junction. In contrast, the majority of occludin (71 +/- 3%) diffuses rapidly within the tight junction with a diffusion constant of 0.011 microm(2)s(-1). Zonula occludens-1 molecules are also highly dynamic in this region, but, rather than diffusing within the plane of the membrane, 69 +/- 5% exchange between membrane and intracellular pools in an energy-dependent manner. These data demonstrate that the tight junction undergoes constant remodeling and suggest that this dynamic behavior may contribute to tight junction assembly and regulation.
Figures



References
-
- Anderson, J.M., C.M. Van Itallie, and A.S. Fanning. 2004. Setting up a selective barrier at the apical junction complex. Curr. Opin. Cell Biol. 16:140–145. - PubMed
-
- Balda, M.S., J.M. Anderson, and K. Matter. 1996. The SH3 domain of the tight junction protein ZO-1 binds to a serine protein kinase that phosphorylates a region C-terminal to this domain. FEBS Lett. 399:326–332. - PubMed
-
- Bazzoni, G., O.M. Martinez-Estrada, F. Orsenigo, M. Cordenonsi, S. Citi, and E. Dejana. 2000. Interaction of junctional adhesion molecule with the tight junction components ZO-1, cingulin, and occludin. J. Biol. Chem. 275:20520–20526. - PubMed
-
- Beatch, M., L.A. Jesaitis, W.J. Gallin, D.A. Goodenough, and B.R. Stevenson. 1996. The tight junction protein ZO-2 contains three PDZ (PSD-95/Discs-Large/ZO-1) domains and an alternatively spliced region. J. Biol. Chem. 271:25723–25726. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

