R5X4 viruses are evolutionary, functional, and antigenic intermediates in the pathway of a simian-human immunodeficiency virus coreceptor switch
- PMID: 18480460
- PMCID: PMC2446975
- DOI: 10.1128/JVI.00570-08
R5X4 viruses are evolutionary, functional, and antigenic intermediates in the pathway of a simian-human immunodeficiency virus coreceptor switch
Abstract
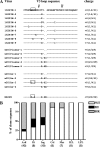
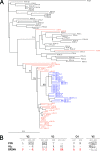
To examine the pathway of the coreceptor switching of CCR5-using (R5) virus to CXCR4-using (X4) virus in simian-human immunodeficiency virus SHIV(SF162P3N)-infected rhesus macaque BR24, analysis was performed on variants present at 20 weeks postinfection, the time when the signature gp120 V3 loop sequence of the X4 switch variant was first detected by PCR. Unexpectedly, circulating and tissue variants with His/Ile instead of the signature X4 V3 His/Arg insertions predominated at this time point. Phylogenetic analysis of the sequences of the C2 conserved region to the V5 variable loop of the envelope (Env) protein showed that viruses bearing HI insertions represented evolutionary intermediates between the parental SHIV(SF162P3N) and the final X4 HR switch variant. Functional analyses demonstrated that the HI variants were phenotypic intermediates as well, capable of using both CCR5 and CXCR4 for entry. However, the R5X4 intermediate virus entered CCR5-expressing target cells less efficiently than the parental R5 strain and was more sensitive to both CCR5 and CXCR4 inhibitors than either the parental R5 or the final X4 virus. It was also more sensitive than the parental R5 virus to antibody neutralization, especially to agents directed against the CD4 binding site, but not as sensitive as the late X4 virus. Significantly, the V3 loop sequence that determined CXCR4 use also conferred soluble CD4 neutralization sensitivity. Collectively, the data illustrate that, similar to human immunodeficiency virus type 1 (HIV-1) infection in individuals, the evolution from CCR5 to CXCR4 usage in BR24 transitions through an intermediate phase with reduced virus entry and coreceptor usage efficiencies. The data further support a model linking an open envelope gp120 conformation, better CD4 binding, and expansion to CXCR4 usage.
Figures
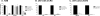
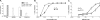
Similar articles
-
Identification of interdependent variables that influence coreceptor switch in R5 SHIV(SF162P3N)-infected macaques.Retrovirology. 2012 Dec 13;9:106. doi: 10.1186/1742-4690-9-106. Retrovirology. 2012. PMID: 23237529 Free PMC article.
-
Fitness disadvantage of transitional intermediates contributes to dynamic change in the infecting-virus population during coreceptor switch in R5 simian/human immunodeficiency virus-infected macaques.J Virol. 2010 Dec;84(24):12862-71. doi: 10.1128/JVI.01478-10. Epub 2010 Oct 13. J Virol. 2010. PMID: 20943985 Free PMC article.
-
Adoption of an "open" envelope conformation facilitating CD4 binding and structural remodeling precedes coreceptor switch in R5 SHIV-infected macaques.PLoS One. 2011;6(7):e21350. doi: 10.1371/journal.pone.0021350. Epub 2011 Jul 8. PLoS One. 2011. PMID: 21760891 Free PMC article.
-
Coreceptor switch in infection of nonhuman primates.Curr HIV Res. 2009 Jan;7(1):30-8. doi: 10.2174/157016209787048500. Curr HIV Res. 2009. PMID: 19149552 Review.
-
Chemokine receptor utilization and macrophage signaling by human immunodeficiency virus type 1 gp120: Implications for neuropathogenesis.J Neurovirol. 2004;10 Suppl 1:91-6. doi: 10.1080/753312758. J Neurovirol. 2004. PMID: 14982745 Review.
Cited by
-
Molecularly cloned SHIV-CN97001: a replication-competent, R5 simian/human immunodeficiency virus containing env of a primary Chinese HIV-1 clade C isolate.J Med Primatol. 2011 Dec;40(6):427-36. doi: 10.1111/j.1600-0684.2011.00497.x. Epub 2011 Sep 6. J Med Primatol. 2011. PMID: 21895680 Free PMC article.
-
SHIV-1157i and passaged progeny viruses encoding R5 HIV-1 clade C env cause AIDS in rhesus monkeys.Retrovirology. 2008 Oct 17;5:94. doi: 10.1186/1742-4690-5-94. Retrovirology. 2008. PMID: 18928523 Free PMC article.
-
Pathogenic consequences of vaginal infection with CCR5-tropic simian-human immunodeficiency virus SHIVSF162P3N.J Virol. 2012 Sep;86(17):9432-42. doi: 10.1128/JVI.00852-12. Epub 2012 Jun 27. J Virol. 2012. PMID: 22740397 Free PMC article.
-
Modulation of the virus-receptor interaction by mutations in the V5 loop of feline immunodeficiency virus (FIV) following in vivo escape from neutralising antibody.Retrovirology. 2010 Apr 26;7:38. doi: 10.1186/1742-4690-7-38. Retrovirology. 2010. PMID: 20420700 Free PMC article.
-
Understanding the HIV coreceptor switch from a dynamical perspective.BMC Evol Biol. 2009 Nov 30;9:274. doi: 10.1186/1471-2148-9-274. BMC Evol Biol. 2009. PMID: 19948048 Free PMC article.
References
-
- Baik, S. S., R. W. Doms, and B. J. Doranz. 1999. HIV and SIV gp120 binding does not predict coreceptor function. Virology 259267-273. - PubMed
-
- Berger, E. A., R. W. Doms, E. M. Fenyo, B. T. Korber, D. R. Littman, J. P. Moore, Q. J. Sattentau, H. Schuitemaker, J. Sodroski, and R. A. Weiss. 1998. A new classification for HIV-1. Nature 391240. - PubMed
-
- Brenchley, J. M., T. W. Schacker, L. E. Ruff, D. A. Price, J. H. Taylor, G. J. Beilman, P. L. Nguyen, A. Khoruts, M. Larson, A. T. Haase, and D. C. Douek. 2004. CD4+ T cell depletion during all stages of HIV disease occurs predominantly in the gastrointestinal tract. J. Exp. Med. 200749-759. - PMC - PubMed
-
- Brumme, Z. L., J. Goodrich, H. B. Mayer, C. J. Brumme, B. M. Henrick, B. Wynhoven, J. J. Asselin, P. K. Cheung, R. S. Hogg, J. S. Montaner, and P. R. Harrigan. 2005. Molecular and clinical epidemiology of CXCR4-using HIV-1 in a large population of antiretroviral-naive individuals. J. Infect. Dis. 192466-474. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

