PP2B and PP1alpha cooperatively disrupt 7SK snRNP to release P-TEFb for transcription in response to Ca2+ signaling
- PMID: 18483222
- PMCID: PMC2377190
- DOI: 10.1101/gad.1636008
PP2B and PP1alpha cooperatively disrupt 7SK snRNP to release P-TEFb for transcription in response to Ca2+ signaling
Abstract
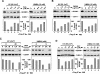
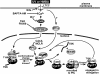
The positive transcription elongation factor b (P-TEFb), consisting of Cdk9 and cyclin T, stimulates RNA polymerase II elongation and cotranscriptional pre-mRNA processing. To accommodate different growth conditions and transcriptional demands, a reservoir of P-TEFb is kept in an inactive state in the multisubunit 7SK snRNP. Under certain stress or disease conditions, P-TEFb is released to activate transcription, although the signaling pathway(s) that controls this is largely unknown. Here, through analyzing the UV- or hexamethylene bisacetamide (HMBA)-induced release of P-TEFb from 7SK snRNP, an essential role for the calcium ion (Ca2+)-calmodulin-protein phosphatase 2B (PP2B) signaling pathway is revealed. However, Ca2+ signaling alone is insufficient, and PP2B must act sequentially and cooperatively with protein phosphatase 1alpha (PP1alpha) to disrupt 7SK snRNP. Activated by UV/HMBA and facilitated by a PP2B-induced conformational change in 7SK snRNP, PP1alpha releases P-TEFb through dephosphorylating phospho-Thr186 in the Cdk9 T-loop. This event is also necessary for the subsequent recruitment of P-TEFb by the bromodomain protein Brd4 to the preinitiation complex, where Cdk9 remains unphosphorylated and inactive until after the synthesis of a short RNA. Thus, through cooperatively dephosphorylating Cdk9 in response to Ca2+ signaling, PP2B and PP1alpha alter the P-TEFb functional equilibrium through releasing P-TEFb from 7SK snRNP for transcription.
Figures





References
-
- Chao S.H., Price D.H. Flavopiridol inactivates P-TEFb and blocks most RNA polymerase II transcription in vivo. J. Biol. Chem. 2001;276:31793–31799. - PubMed
-
- Chen R., Yang Z., Zhou Q. Phosphorylated positive transcription elongation factor b (P-TEFb) is tagged for inhibition through association with 7SK snRNA. J. Biol. Chem. 2004;279:4153–4160. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
