Were there any "misassignments" among iron-sulfur clusters N4, N5 and N6b in NADH-quinone oxidoreductase (complex I)?
- PMID: 18486592
- PMCID: PMC2724968
- DOI: 10.1016/j.bbabio.2008.04.032
Were there any "misassignments" among iron-sulfur clusters N4, N5 and N6b in NADH-quinone oxidoreductase (complex I)?
Abstract
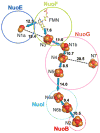
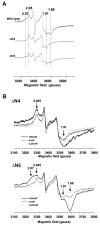
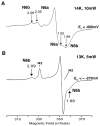
NADH-quinone oxidoreductase (complex I) in bovine heart mitochondria has a molecular weight of approximately 1 million Da composed of 45 distinct subunits. It is the largest energy transducing complex so far known. Bacterial complex I is simpler and smaller, but the essential redox components and the basic mechanisms of electron and proton translocation are the same. Over the past three decades, Ohnishi et al. have pursued extensive EPR studies near liquid helium temperatures and characterized most of the iron-sulfur clusters in complex I. Recently, Yakovlev et al. [G. Yakovlev, T. Reda, J. Hirst, Reevaluating the relationship between EPR spectra and enzyme structure for the iron-sulfur clusters in NADH:quinone oxidoreductase, Proc. Natl. Acad. Sci. U. S. A. 104 (2007) 12720-12725] challenged Ohnishi's group by claiming that there were EPR "misassignments" among clusters N4, N5 and N6b (in order to prevent confusion, we used current consensus nomenclature, as the nickname). They claimed that we misassigned EPR signals arising from cluster N5 to cluster N4, and signals from cluster N6b to cluster N4. They also proposed that cluster N5 has (4Cys)-ligands. Based on the accumulated historical data and recent results of our site-specific mutagenesis experiments, we confirmed that cluster N5 has (1His+3Cys)-ligands as we had predicted. We revealed that E. coli cluster N5 signals could be clearly detected at the sample temperature around 3 K with microwave power higher than 5 mW. Thus Hirst's group could not detect N5 signals under any of their EPR conditions, reported in their PNAS paper. It seems that they misassigned the signals from cluster N4 to N5. As to the claim of "misassignment" between clusters N4 and N6b, that was not a possibility because our mutagenesis systems did not contain cluster N6b. Therefore, we believe that we have not made any "misassignment" in our work.
Figures
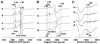
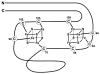
Similar articles
-
Iron-sulfur cluster N5 is coordinated by an HXXXCXXCXXXXXC motif in the NuoG subunit of Escherichia coli NADH:quinone oxidoreductase (complex I).J Biol Chem. 2008 Sep 19;283(38):25979-87. doi: 10.1074/jbc.M804015200. Epub 2008 Jul 4. J Biol Chem. 2008. PMID: 18603533 Free PMC article.
-
Characterization of the iron-sulfur cluster N7 (N1c) in the subunit NuoG of the proton-translocating NADH-quinone oxidoreductase from Escherichia coli.J Biol Chem. 2005 Jan 7;280(1):301-7. doi: 10.1074/jbc.M410377200. Epub 2004 Nov 1. J Biol Chem. 2005. PMID: 15520003
-
Reevaluating the relationship between EPR spectra and enzyme structure for the iron sulfur clusters in NADH:quinone oxidoreductase.Proc Natl Acad Sci U S A. 2007 Jul 31;104(31):12720-5. doi: 10.1073/pnas.0705593104. Epub 2007 Jul 19. Proc Natl Acad Sci U S A. 2007. PMID: 17640900 Free PMC article.
-
Iron-sulfur clusters/semiquinones in complex I.Biochim Biophys Acta. 1998 May 6;1364(2):186-206. doi: 10.1016/s0005-2728(98)00027-9. Biochim Biophys Acta. 1998. PMID: 9593887 Review.
-
A possible role for iron-sulfur cluster N2 in proton translocation by the NADH: ubiquinone oxidoreductase (complex I).J Mol Microbiol Biotechnol. 2005;10(2-4):208-22. doi: 10.1159/000091566. J Mol Microbiol Biotechnol. 2005. PMID: 16645316 Review.
Cited by
-
Architecture of complex I and its implications for electron transfer and proton pumping.Biochim Biophys Acta. 2009 Jun;1787(6):574-83. doi: 10.1016/j.bbabio.2009.01.012. Epub 2009 Feb 7. Biochim Biophys Acta. 2009. PMID: 19366614 Free PMC article. Review.
-
Mössbauer spectroscopy on respiratory complex I: the iron-sulfur cluster ensemble in the NADH-reduced enzyme is partially oxidized.Biochemistry. 2012 Jan 10;51(1):149-58. doi: 10.1021/bi201644x. Epub 2011 Dec 9. Biochemistry. 2012. PMID: 22122402 Free PMC article.
-
Essential regions in the membrane domain of bacterial complex I (NDH-1): the machinery for proton translocation.J Bioenerg Biomembr. 2014 Aug;46(4):279-87. doi: 10.1007/s10863-014-9558-8. Epub 2014 Jun 29. J Bioenerg Biomembr. 2014. PMID: 24973951 Review.
-
The pro-oxidant chromium(VI) inhibits mitochondrial complex I, complex II, and aconitase in the bronchial epithelium: EPR markers for Fe-S proteins.Free Radic Biol Med. 2010 Dec 15;49(12):1903-15. doi: 10.1016/j.freeradbiomed.2010.09.020. Epub 2010 Sep 27. Free Radic Biol Med. 2010. PMID: 20883776 Free PMC article.
-
Semiquinone and cluster N6 signals in His-tagged proton-translocating NADH:ubiquinone oxidoreductase (complex I) from Escherichia coli.J Biol Chem. 2013 May 17;288(20):14310-14319. doi: 10.1074/jbc.M113.467803. Epub 2013 Mar 29. J Biol Chem. 2013. PMID: 23543743 Free PMC article.
References
-
- Walker JE. The NADH:ubiquinone oxidoreductase (complex I) of respiratory chains. Q Rev Biophys. 1992;25:253–324. - PubMed
-
- Ohnishi T. Iron–sulfur clusters/semiquinones in Complex I. Biochim Biophys Acta. 1998;1364:186–206. - PubMed
-
- Yagi T, Matsuno-Yagi A. The proton-translocating NADH-quinone oxidoreductase in the respiratory chain: the secret unlocked. Biochemistry. 2003;42:2266–2274. - PubMed
-
- Sazanov LA, Hinchliffe P. Structure of the hydrophilic domain of respiratory complex I from Thernus thermophilus. Science. 2006;311:1430–1436. - PubMed
-
- Sazanov LA. Respiratory complex I: mechanistic and structural insights provided by the crystal structure of the hydrophilic domain. Biochemistry. 2007;46:2275–2288. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

