Dual effects of miR156-targeted SPL genes and CYP78A5/KLUH on plastochron length and organ size in Arabidopsis thaliana
- PMID: 18492871
- PMCID: PMC2438454
- DOI: 10.1105/tpc.108.058180
Dual effects of miR156-targeted SPL genes and CYP78A5/KLUH on plastochron length and organ size in Arabidopsis thaliana
Abstract
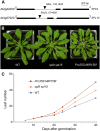
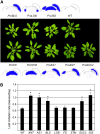
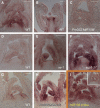
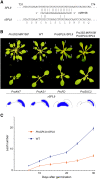
Leaves of flowering plants are produced from the shoot apical meristem at regular intervals, with the time that elapses between the formation of two successive leaf primordia defining the plastochron. We have identified two genetic axes affecting plastochron length in Arabidopsis thaliana. One involves microRNA156 (miR156), which targets a series of SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) genes. In situ hybridization studies and misexpression experiments demonstrate that miR156 is a quantitative, rather than spatial, modulator of SPL expression in leaf primordia and that SPL activity nonautonomously inhibits initiation of new leaves at the shoot apical meristem. The second axis is exemplified by a redundantly acting pair of cytochrome P450 genes, CYP78A5/KLUH and CYP78A7, which are likely orthologs of PLASTOCHRON1 of rice (Oryza sativa). Inactivation of CYP78A5, which is expressed at the periphery of the shoot apical meristem, accelerates the leaf initiation rate, whereas cyp78a5 cyp78a7 double mutants often die as embryos with supernumerary cotyledon primordia. The effects of both miR156-targeted SPL genes and CYP78A5 on organ size are correlated with changes in plastochron length, suggesting a potential compensatory mechanism that links the rate at which leaves are produced to final leaf size.
Figures


Similar articles
-
ALTERED MERISTEM PROGRAM1 regulates leaf identity independently of miR156-mediated translational repression.Development. 2020 Apr 27;147(8):dev186874. doi: 10.1242/dev.186874. Development. 2020. PMID: 32198155 Free PMC article.
-
Gradual increase of miR156 regulates temporal expression changes of numerous genes during leaf development in rice.Plant Physiol. 2012 Mar;158(3):1382-94. doi: 10.1104/pp.111.190488. Epub 2012 Jan 23. Plant Physiol. 2012. PMID: 22271747 Free PMC article.
-
Strigolactones (SLs) modulate the plastochron by regulating KLUH (KLU) transcript abundance in Arabidopsis.New Phytol. 2021 Dec;232(5):1909-1916. doi: 10.1111/nph.17725. Epub 2021 Sep 24. New Phytol. 2021. PMID: 34498760
-
A Regulatory Network for miR156-SPL Module in Arabidopsis thaliana.Int J Mol Sci. 2019 Dec 6;20(24):6166. doi: 10.3390/ijms20246166. Int J Mol Sci. 2019. PMID: 31817723 Free PMC article. Review.
-
From genes to networks: The genetic control of leaf development.J Integr Plant Biol. 2021 Jul;63(7):1181-1196. doi: 10.1111/jipb.13084. Epub 2021 Apr 14. J Integr Plant Biol. 2021. PMID: 33615731 Review.
Cited by
-
Establishment of the Embryonic Shoot Meristem Involves Activation of Two Classes of Genes with Opposing Functions for Meristem Activities.Int J Mol Sci. 2020 Aug 15;21(16):5864. doi: 10.3390/ijms21165864. Int J Mol Sci. 2020. PMID: 32824181 Free PMC article.
-
Control of cell proliferation in Arabidopsis thaliana by microRNA miR396.Development. 2010 Jan;137(1):103-12. doi: 10.1242/dev.043067. Development. 2010. PMID: 20023165 Free PMC article.
-
Anisotropic cell growth at the leaf base promotes age-related changes in leaf shape in Arabidopsis thaliana.Plant Cell. 2023 Apr 20;35(5):1386-1407. doi: 10.1093/plcell/koad031. Plant Cell. 2023. PMID: 36748203 Free PMC article.
-
Diurnal Regulation of Plant Epidermal Wax Synthesis through Antagonistic Roles of the Transcription Factors SPL9 and DEWAX.Plant Cell. 2019 Nov;31(11):2711-2733. doi: 10.1105/tpc.19.00233. Epub 2019 Sep 4. Plant Cell. 2019. PMID: 31484683 Free PMC article.
-
Contribution of Omics and Systems Biology to Plant Biotechnology.Adv Exp Med Biol. 2021;1346:171-188. doi: 10.1007/978-3-030-80352-0_10. Adv Exp Med Biol. 2021. PMID: 35113402 Review.
References
-
- Abe, M., Kobayashi, Y., Yamamoto, S., Daimon, Y., Yamaguchi, A., Ikeda, Y., Ichinoki, H., Notaguchi, M., Goto, K., and Araki, T. (2005). FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 309 1052–1056. - PubMed
-
- Adenot, X., Elmayan, T., Lauressergues, D., Boutet, S., Bouche, N., Gasciolli, V., and Vaucheret, H. (2006). DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr. Biol. 16 927–932. - PubMed
-
- Alonso, J.M., et al. (2003). Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301 653–657. - PubMed
-
- Anastasiou, E., Kenz, S., Gerstung, M., MacLean, D., Timmer, J., Fleck, C., and Lenhard, M. (2007). Control of plant organ size by KLUH/CYP78A5-dependent intercellular signaling. Dev. Cell 13 843–856. - PubMed
-
- Anderson, G.H., Alvarez, N.D.G., Gilman, C., Jeffares, D.C., Trainor, V.C.W., Hanson, M.R., and Veit, B. (2004). Diversification of genes encoding Mei2-Like RNA binding proteins in plants. Plant Mol. Biol. 54 653–670. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

