Hydrogen exchange of monomeric alpha-synuclein shows unfolded structure persists at physiological temperature and is independent of molecular crowding in Escherichia coli
- PMID: 18493022
- PMCID: PMC2492816
- DOI: 10.1110/ps.033803.107
Hydrogen exchange of monomeric alpha-synuclein shows unfolded structure persists at physiological temperature and is independent of molecular crowding in Escherichia coli
Abstract
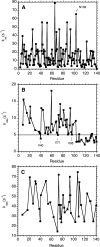
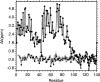
Amide proton NMR signals from the N-terminal domain of monomeric alpha-synuclein (alphaS) are lost when the sample temperature is raised from 10 degrees C to 35 degrees C at pH 7.4. Although the temperature-induced effects have been attributed to conformational exchange caused by an increase in alpha-helix structure, we show that the loss of signals is due to fast amide proton exchange. At low ionic strength, hydrogen exchange rates are faster for the N-terminal segment of alphaS than for the acidic C-terminal domain. When the salt concentration is raised to 300 mM, exchange rates increase throughout the protein and become similar for the N- and C-terminal domains. This indicates that the enhanced protection of amide protons from the C-terminal domain at low salt is electrostatic in nature. Calpha chemical shift data point to <10% residual alpha-helix structure at 10 degrees C and 35 degrees C. Conformational exchange contributions to R2 are negligible at both temperatures. In contrast to the situation in vitro, the majority of amide protons are observed at 37 degrees C in 1H-15N HSQC spectra of alphaS encapsulated within living Escherichia coli cells. Our finding that temperature effects on alphaS NMR spectra can be explained by hydrogen exchange obviates the need to invoke special cellular factors. The retention of signals is likely due to slowed hydrogen exchange caused by the lowered intracellular pH of high-density E. coli cultures. Taken together, our results emphasize that alphaS remains predominantly unfolded at physiological temperature and pH-an important conclusion for mechanistic models of the association of alphaS with membranes and fibrils.
Figures



Similar articles
-
Isolation of recombinant tetrameric N-acetylated α-synuclein.Protein Expr Purif. 2018 Dec;152:146-154. doi: 10.1016/j.pep.2018.07.008. Epub 2018 Jul 21. Protein Expr Purif. 2018. PMID: 30041032
-
Distinct residual and disordered structures of alpha-synuclein analyzed by amide-proton exchange and NMR signal intensity.Biochim Biophys Acta Proteins Proteom. 2020 Sep;1868(9):140464. doi: 10.1016/j.bbapap.2020.140464. Epub 2020 Jun 2. Biochim Biophys Acta Proteins Proteom. 2020. PMID: 32497661
-
NMR determination of pKa values in α-synuclein.Protein Sci. 2011 Feb;20(2):256-69. doi: 10.1002/pro.556. Epub 2010 Dec 13. Protein Sci. 2011. PMID: 21280118 Free PMC article.
-
α-Synuclein and neuronal membranes: Conformational flexibilities in health and disease.Chem Phys Lipids. 2021 Mar;235:105034. doi: 10.1016/j.chemphyslip.2020.105034. Epub 2021 Jan 9. Chem Phys Lipids. 2021. PMID: 33434528 Review.
-
Disruptive membrane interactions of alpha-synuclein aggregates.Biochim Biophys Acta Proteins Proteom. 2019 May;1867(5):468-482. doi: 10.1016/j.bbapap.2018.10.006. Epub 2018 Oct 11. Biochim Biophys Acta Proteins Proteom. 2019. PMID: 30315896 Review.
Cited by
-
The Residual Structure of Unfolded Proteins was Elucidated from the Standard Deviation of NMR Intensity Differences.Protein Pept Lett. 2023;30(2):103-107. doi: 10.2174/0929866530666230104140830. Protein Pept Lett. 2023. PMID: 36600624 Free PMC article.
-
The molecular mechanism of nuclear transport revealed by atomic-scale measurements.Elife. 2015 Sep 15;4:e10027. doi: 10.7554/eLife.10027. Elife. 2015. PMID: 26371551 Free PMC article.
-
Contextual Role of a Salt Bridge in the Phage P22 Coat Protein I-Domain.J Biol Chem. 2016 May 20;291(21):11359-72. doi: 10.1074/jbc.M116.716910. Epub 2016 Mar 22. J Biol Chem. 2016. PMID: 27006399 Free PMC article.
-
CP-HISQC: a better version of HSQC experiment for intrinsically disordered proteins under physiological conditions.J Biomol NMR. 2014 Mar;58(3):175-92. doi: 10.1007/s10858-014-9815-5. Epub 2014 Feb 5. J Biomol NMR. 2014. PMID: 24496557
-
Physicochemical properties of cells and their effects on intrinsically disordered proteins (IDPs).Chem Rev. 2014 Jul 9;114(13):6661-714. doi: 10.1021/cr400695p. Epub 2014 Jun 5. Chem Rev. 2014. PMID: 24901537 Free PMC article. Review. No abstract available.
References
-
- Alexandrescu, A.T., Abeygunawardana, C., Shortle, D. Structure and dynamics of a denatured 131-residue fragment of staphylococcal nuclease: A heteronuclear NMR study. Biochemistry. 1994;33:1063–1072. - PubMed
-
- Alexandrescu, A.T., Jahnke, W., Wiltscheck, R., Blommers, M.J. Accretion of structure in staphylococcal nuclease: A 15N NMR relaxation study. J. Mol. Biol. 1996;260:570–587. - PubMed
-
- Basu, A., Glew, R.H. Characterization of the activation of rat liver β-glucosidase by sialosylgangliotetraosylceramide. J. Biol. Chem. 1985;260:13067–13073. - PubMed
-
- Bermel, W., Bertini, I., Felli, I.C., Lee, Y.M., Luchinat, C., Pierattelli, R. Protonless NMR experiments for sequence-specific assignment of backbone nuclei in unfolded proteins. J. Am. Chem. Soc. 2006;128:3918–3919. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

