Barren inflorescence1 functions in organogenesis during vegetative and inflorescence development in maize
- PMID: 18493061
- PMCID: PMC2390617
- DOI: 10.1534/genetics.107.084079
Barren inflorescence1 functions in organogenesis during vegetative and inflorescence development in maize
Abstract
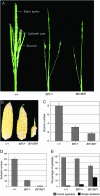
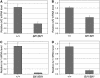
Maize (Zea mays) has a highly branched inflorescence due to the production of different types of axillary meristems. Characterization of the barren inflorescence class of mutants has led to the discovery of genes required for axillary meristem initiation in the inflorescence. Previous studies showed that barren inflorescence2 (bif2) encodes a serine/threonine protein kinase that regulates auxin transport, and barren stalk1 (ba1) encodes a basic helix-loop-helix transcription factor that acts downstream of auxin transport. Here, we characterize Barren inflorescence1 (Bif1), a classical semidominant mutation of maize. Developmental, histological, and genetic analyses show that Bif1 mutants are defective in the initiation of all axillary meristems in the inflorescence. Real time RT-PCR experiments show that both bif2 and ba1 are expressed at lower levels in Bif1 mutants. Double-mutant analyses demonstrate that Bif1 exhibits an epistatic interaction with ba1 and a synergistic interaction with bif2. The dramatic phenotypic enhancement observed in Bif1; bif2 double mutants implies that bif1 plays an overlapping role with bif2 in the initiation of lateral organs during vegetative development. The phenotypic resemblance of Bif1 to bif2 mutants and the reduction of auxin transport in Bif1 mutants suggest that bif1 functions as a regulator of auxin transport in maize.
Figures



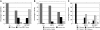



References
-
- Alleman, M., L. Sidorenko, K. McGinnis, V. Seshadri, J. E. Dorweiler et al., 2006. An RNA-dependent RNA polymerase is required for paramutation in maize. Nature 442 295–298. - PubMed
-
- Benjamins, R., A. Quint, D. Weijers, P. Hooykaas and R. Offringa, 2001. The PINOID protein kinase regulates organ development in Arabidopsis by enhancing polar auxin transport. Development 128 4057–4067. - PubMed
-
- Benkova, E., M. Michniewicz, M. Sauer, T. Teichmann, D. Seifertova et al., 2003. Local, efflux-dependent auxin gradients as a common module for plant organ formation. Cell 115 591–602. - PubMed
-
- Bennett, S. R. M., J. Alvarez, G. Bossinger and D. R. Smyth, 1995. Morphogenesis in pinoid mutants of Arabidopsis thaliana. Plant J. 8 505–520.
-
- Bennett, T., T. Sieberer, B. Willett, J. Booker, C. Luschnig et al., 2006. The Arabidopsis MAX pathway controls shoot branching by regulating auxin transport. Curr. Biol. 16 553–563. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

