Development of multiplex PCR-ligase detection reaction assay for detection of West Nile virus
- PMID: 18495862
- PMCID: PMC2446923
- DOI: 10.1128/JCM.02335-07
Development of multiplex PCR-ligase detection reaction assay for detection of West Nile virus
Abstract
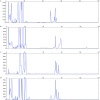
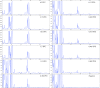
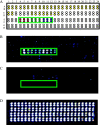
We have developed a novel multiplex reverse transcription-PCR ligase detection reaction (RT-PCR/LDR) assay for the detection of West Nile virus (WNV) in both clinical and mosquito pool samples. The method relies on the amplification of three different genomic regions, one in the coding sequence of nonstructural protein NS2a and two in nonstructural protein NS5, to minimize the risk of detection failure due to genetic variation. The sensitivity of the PCR is complemented by the high specificity of the LDR step, and the detection of the LDR products can be achieved with capillary electrophoresis (CE) or a universal DNA microarray. We evaluated the limit of detection by both one-step and two-step multiplex RT-PCR/LDR/CE approaches, which reached, respectively, 0.005 and 0.017 PFU. The assay demonstrated 99% sensitivity when mosquito pool samples were tested and 100% sensitivity with clinical samples when the one-step approach was used. The broad strain coverage was confirmed by testing 34 WNV isolates belonging to lineages 1 and 2, and the high specificity of the assay was determined by testing other flaviviruses, as well as negative mosquito pool and clinical samples. In summary, the multiplex RT-PCR/LDR assay could represent a valuable complement to WNV serological diagnosis, especially in early symptomatic patients. In addition, the multiplexing capacity of the technique, which can be coupled to universal DNA microarray detection, makes it an amenable tool to develop a more comprehensive assay for viral pathogens.
Figures

Similar articles
-
Simultaneous detection of West Nile and Japanese encephalitis virus RNA by duplex TaqMan RT-PCR.J Virol Methods. 2013 Nov;193(2):554-7. doi: 10.1016/j.jviromet.2013.07.025. Epub 2013 Jul 25. J Virol Methods. 2013. PMID: 23892127
-
A novel quantitative multiplex real-time RT-PCR for the simultaneous detection and differentiation of West Nile virus lineages 1 and 2, and of Usutu virus.J Virol Methods. 2013 May;189(2):321-7. doi: 10.1016/j.jviromet.2013.02.019. Epub 2013 Mar 7. J Virol Methods. 2013. PMID: 23499258
-
Real time PCR assay for detection of all known lineages of West Nile virus.J Virol Methods. 2016 Oct;236:266-270. doi: 10.1016/j.jviromet.2016.07.026. Epub 2016 Jul 30. J Virol Methods. 2016. PMID: 27481597
-
Highly sensitive TaqMan RT-PCR assay for detection and quantification of both lineages of West Nile virus RNA.J Clin Virol. 2006 Jul;36(3):177-82. doi: 10.1016/j.jcv.2006.02.008. Epub 2006 May 3. J Clin Virol. 2006. PMID: 16675298
-
Vector surveillance for West Nile virus.Ann N Y Acad Sci. 2001 Dec;951:74-83. doi: 10.1111/j.1749-6632.2001.tb02686.x. Ann N Y Acad Sci. 2001. PMID: 11797806 Review.
Cited by
-
Analysis of the genetic effects of prolactin gene polymorphisms on chicken egg production.Mol Biol Rep. 2013 Jan;40(1):289-94. doi: 10.1007/s11033-012-2060-7. Epub 2012 Nov 25. Mol Biol Rep. 2013. PMID: 23184001
-
A multiplex PCR/LDR assay for simultaneous detection and identification of the NIAID category B bacterial food and water-borne pathogens.Diagn Microbiol Infect Dis. 2014 Jun;79(2):135-40. doi: 10.1016/j.diagmicrobio.2014.02.022. Epub 2014 Mar 12. Diagn Microbiol Infect Dis. 2014. PMID: 24709368 Free PMC article.
-
Advances in ligase chain reaction and ligation-based amplifications for genotyping assays: Detection and applications.Mutat Res Rev Mutat Res. 2017 Jul;773:66-90. doi: 10.1016/j.mrrev.2017.05.001. Epub 2017 May 2. Mutat Res Rev Mutat Res. 2017. PMID: 28927538 Free PMC article. Review.
-
Multiplex PCR-ligation detection reaction assay for simultaneous detection of drug resistance and toxin genes from Staphylococcus aureus, Enterococcus faecalis, and Enterococcus faecium.J Clin Microbiol. 2010 Jan;48(1):277-80. doi: 10.1128/JCM.01411-09. Epub 2009 Oct 28. J Clin Microbiol. 2010. PMID: 19864481 Free PMC article.
-
Development of a flow-through [corrected] microarray based reverse transcriptase multiplex ligation-dependent probe amplification assay for the detection of European Bunyaviruses. [corrected].Mol Biotechnol. 2011 Oct;49(2):176-86. doi: 10.1007/s12033-011-9389-3. Mol Biotechnol. 2011. PMID: 21390485 Free PMC article.
References
-
- Barany, F., and D. H. Gelfand. 1991. Cloning, overexpression and nucleotide sequence of a thermostable DNA ligase-encoding gene. Gene 1091-11. - PubMed
-
- Beasley, D. W., C. T. Davis, H. Guzman, D. L. Vanlandingham, A. P. Travassos da Rosa, R. E. Parsons, S. Higgs, R. B. Tesh, and A. D. Barrett. 2003. Limited evolution of West Nile virus has occurred during its southwesterly spread in the United States. Virology 309190-195. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

