Extending the breadth of metabolite profiling by gas chromatography coupled to mass spectrometry
- PMID: 18497891
- PMCID: PMC2394193
- DOI: 10.1016/j.trac.2008.01.007
Extending the breadth of metabolite profiling by gas chromatography coupled to mass spectrometry
Abstract
Gas chromatography coupled to mass spectrometry (GC-MS) is one of the most frequently used tools for profiling primary metabolites. Instruments are mature enough to run large sequences of samples; novel advancements increase the breadth of compounds that can be analyzed, and improved algorithms and databases are employed to capture and utilize biologically relevant information. Around half the published reports on metabolite profiling by GC-MS focus on biological problems rather than on methodological advances. Applications span from comprehensive analysis of volatiles to assessment of metabolic fluxes for bioengineering. Method improvements emphasize extraction procedures, evaluations of quality control of GC-MS in comparison to other techniques and approaches to data processing. Two major challenges remain: rapid annotation of unknown peaks; and, integration of biological background knowledge aiding data interpretation.
Figures
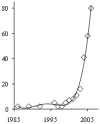



References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
