Novel role for SLPI in MOG-induced EAE revealed by spinal cord expression analysis
- PMID: 18501024
- PMCID: PMC2438345
- DOI: 10.1186/1742-2094-5-20
Novel role for SLPI in MOG-induced EAE revealed by spinal cord expression analysis
Abstract
Background: Experimental autoimmune encephalomyelitis (EAE) induced by myelin oligodendrocyte protein (MOG) in female Dark Agouti (DA) rats is a chronic demyelinating animal model of multiple sclerosis (MS). To identify new candidate molecules involved in the evolution or repair of EAE-lesions we used Affymetrix oligonucleotide microarrays to compare the spinal cord transcriptome at the peak of EAE, during remission and at the first relapse with healthy DA rats.
Methods: Untreated DA rats and DA rats immunised with MOG protein were sacrificed at defined time points. Total RNA was isolated from spinal cord tissue and used for hybridization of Affymetrix rat genome arrays RG U34 A-C. Selected expression values were confirmed by RealTime PCR. Adult neural stem cells were incubated with recombinant secretory leukocyte protease inhibitor (SLPI). Proliferation was assessed by BrdU incorporation, cyclin D1 and HES1 expression by RealTime PCR, cell differentiation by immunofluorescence analysis and I kappa B alpha degradation by Western blot.
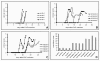
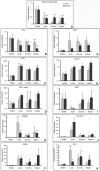
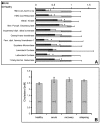
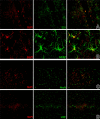
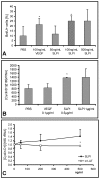
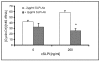
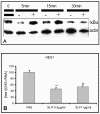
Results: Among approximately 26,000 transcripts studied more than 1,100 were differentially regulated. Focussing on functional themes, we noticed a sustained downregulation of most of the transcripts of the cholesterol biosynthesis pathway. Furthermore, we found new candidate genes possibly contributing to regenerative processes in the spinal cord. Twelve transcripts were solely upregulated in the recovery phase, including genes not previously associated with repair processes. Expression of SLPI was upregulated more than hundredfold during EAE attack. Using immunohistochemistry, SLPI was identified in macrophages, activated microglia, neuronal cells and astrocytes. Incubation of adult neural stem cells (NSC) with recombinant SLPI resulted in an increase of cell proliferation and of differentiation towards oligodendrocytes. These processes were paralleled by an upregulation of the cell-cycle promotor cyclin D1 and a suppression of the cell differentiation regulator HES1. Finally, SLPI prevented the degradation of I kappa B alpha, which may explain the suppression of the cell differentiation inhibitor HES1 suggesting a possible mechanism of oligodendroglial differentiation.
Conclusion: We identified novel features of gene expression in the CNS during EAE, in particular the suppression of genes of cholesterol biosynthesis and a strong upregulation of SLPI, a gene which is for the first time associated with autoimmune inflammation. The capacity of SLPI to increase proliferation of adult NSC and of oligodendroglial differentiation suggests a novel role for SLPI in the promotion of tissue repair, beyond its known functions in the prevention of tissue damages by protease inhibition damage and modulation of inflammatory reactions.
Figures


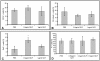
References
-
- Chabas D, Baranzini SE, Mitchell D, Bernard CC, Rittling SR, Denhardt DT, Sobel RA, Lock C, Karpuj M, Pedotti R, Heller R, Oksenberg JR, Steinman L. The influence of the proinflammatory cytokine, osteopontin, on autoimmune demyelinating disease. Science. 2001;294:1731–1735. doi: 10.1126/science.1062960. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

