Sox12 deletion in the mouse reveals nonreciprocal redundancy with the related Sox4 and Sox11 transcription factors
- PMID: 18505825
- PMCID: PMC2493363
- DOI: 10.1128/MCB.00338-08
Sox12 deletion in the mouse reveals nonreciprocal redundancy with the related Sox4 and Sox11 transcription factors
Abstract
The transcription factors Sox4 and Sox11 are important regulators of diverse developmental processes including heart, lung, pancreas, spleen, and B-cell development. Here we have studied the role of the related Sox12 as the third protein of the SoxC group both in vivo and in vitro. Despite widespread Sox12 expression during embryonic development, Sox12-deficient mice developed surprisingly normally, so that they were born alive, showed no gross phenotypic abnormalities, and were fertile in both sexes. Comparison with the related Sox4 and Sox11 revealed extensive overlap in the embryonic expression pattern but more uniform expression levels for Sox12, without sites of particularly high expression. All three Sox proteins furthermore exhibited comparable DNA-binding characteristics and functioned as transcriptional activators. Sox12 was, however, a relatively weak transactivator in comparison to Sox11. We conclude that Sox4 and Sox11 function redundantly with Sox12 and can compensate its loss during mouse development. Because of differences in expression levels and transactivation rates, however, functional compensation is not reciprocal.
Figures








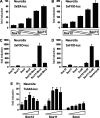

References
-
- Bi, W., J. M. Deng, Z. Zhang, R. R. Behringer, and B. de Crombrugghe. 1999. Sox9 is required for cartilage formation. Nat. Genet. 2285-89. - PubMed
-
- Bowles, J., G. Schepers, and P. Koopman. 2000. Phylogeny of the SOX family of developmental transcription factors based on sequence and structural indicators. Dev. Biol. 227239-255. - PubMed
-
- Cheung, M., M. Abu-Elmagd, H. Clevers, and P. J. Scotting. 2000. Roles of Sox4 in central nervous system development. Mol. Brain Res. 79180-191. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
