Protein and nucleotide damage by glyoxal and methylglyoxal in physiological systems--role in ageing and disease
- PMID: 18533367
- PMCID: PMC2649415
- DOI: 10.1515/dmdi.2008.23.1-2.125
Protein and nucleotide damage by glyoxal and methylglyoxal in physiological systems--role in ageing and disease
Abstract
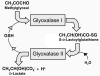
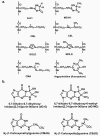
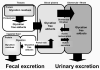
Glycation of proteins, nucleotides and basic phospholipids by glyoxal and methylglyoxal--physiological substrates of glyoxalase 1--is potentially damaging to the proteome, genome and lipidome. Glyoxalase 1 suppresses glycation by these alpha-oxoaldehyde metabolites and thereby represents part of the enzymatic defence against glycation. Albert Szent-Györgyi pioneered and struggled to understand the physiological function of methylglyoxal and the glyoxalase system. We now appreciate that glyoxalase 1 protects against dicarbonyl modifications of the proteome, genome and lipome. Latest research suggests there are functional modifications of this process--implying a role in cell signalling, ageing and disease.
Figures
References
-
- Szent-Gyorgyi A, Egyud LG, McLaughlin JA. Keto-aldehyde and cell division. Science. 1967;155:539–541. - PubMed
-
- Thornalley PJ. Glyoxalase I - structure, function and a critical role in the enzymatic defence against glycation. Biochem Soc Trans. 2003;31:1343–1348. - PubMed
-
- Neuberg C. The destruction of lactic aldehyde and methylglyoxal by animal organs. Biochem Z. 1913;49:502–506.
-
- Cooper RA. Metabolism of methylglyoxal in microorganisms. Ann Rev Microbiol. 1984;38:49–68. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
