Regulation of pyrimidine biosynthetic gene expression in bacteria: repression without repressors
- PMID: 18535147
- PMCID: PMC2415746
- DOI: 10.1128/MMBR.00001-08
Regulation of pyrimidine biosynthetic gene expression in bacteria: repression without repressors
Abstract
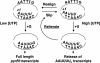
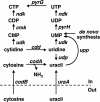
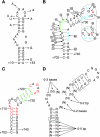
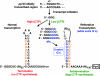
DNA-binding repressor proteins that govern transcription initiation in response to end products generally regulate bacterial biosynthetic genes, but this is rarely true for the pyrimidine biosynthetic (pyr) genes. Instead, bacterial pyr gene regulation generally involves mechanisms that rely only on regulatory sequences embedded in the leader region of the operon, which cause premature transcription termination or translation inhibition in response to nucleotide signals. Studies with Escherichia coli and Bacillus subtilis pyr genes reveal a variety of regulatory mechanisms. Transcription attenuation via UTP-sensitive coupled transcription and translation regulates expression of the pyrBI and pyrE operons in enteric bacteria, whereas nucleotide effects on binding of the PyrR protein to pyr mRNA attenuation sites control pyr operon expression in most gram-positive bacteria. Nucleotide-sensitive reiterative transcription underlies regulation of other pyr genes. With the E. coli pyrBI, carAB, codBA, and upp-uraA operons, UTP-sensitive reiterative transcription within the initially transcribed region (ITR) leads to nonproductive transcription initiation. CTP-sensitive reiterative transcription in the pyrG ITRs of gram-positive bacteria, which involves the addition of G residues, results in the formation of an antiterminator RNA hairpin and suppression of transcription attenuation. Some mechanisms involve regulation of translation rather than transcription. Expression of the pyrC and pyrD operons of enteric bacteria is controlled by nucleotide-sensitive transcription start switching that produces transcripts with different potentials for translation. In Mycobacterium smegmatis and other bacteria, PyrR modulates translation of pyr genes by binding to their ribosome binding site. Evidence supporting these conclusions, generalizations for other bacteria, and prospects for future research are presented.
Figures






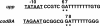



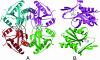
References
-
- Abd-el-Al, A., and J. L. Ingraham. 1969. Control of carbamyl phosphate synthesis in Salmonella typhimurium. J. Biol. Chem. 2444033-4038. - PubMed
-
- Amiott, E. A., and J. A. Jaehning. 2006. Sensitivity of the yeast mitochondrial RNA polymerase to +1 and +2 initiating nucleotides. J. Biol. Chem. 28134982-34988. - PubMed
-
- Andersen, J. T., K. F. Jensen, and P. Poulsen. 1991. Role of transcription pausing in the control of the pyrE attenuator in Escherichia coli. Mol. Microbiol. 5327-333. - PubMed
-
- Andersen, L., M. Kilstrup, and J. Neuhard. 1989. Pyrimidine, purine and nitrogen control of cytosine deaminase synthesis in Escherichia coli K 12. Involvement of the glnLG and purR genes in the regulation of codA expression. Arch. Microbiol. 152115-118. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

