Mechanism of lid closure in the eukaryotic chaperonin TRiC/CCT
- PMID: 18536725
- PMCID: PMC2546500
- DOI: 10.1038/nsmb.1436
Mechanism of lid closure in the eukaryotic chaperonin TRiC/CCT
Abstract
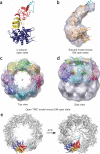
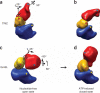
All chaperonins mediate ATP-dependent polypeptide folding by confining substrates within a central chamber. Intriguingly, the eukaryotic chaperonin TRiC (also called CCT) uses a built-in lid to close the chamber, whereas prokaryotic chaperonins use a detachable lid. Here we determine the mechanism of lid closure in TRiC using single-particle cryo-EM and comparative protein modeling. Comparison of TRiC in its open, nucleotide-free, and closed, nucleotide-induced states reveals that the interdomain motions leading to lid closure in TRiC are radically different from those of prokaryotic chaperonins, despite their overall structural similarity. We propose that domain movements in TRiC are coordinated through unique interdomain contacts within each subunit and, further, these contacts are absent in prokaryotic chaperonins. Our findings show how different mechanical switches can evolve from a common structural framework through modification of allosteric networks.
Figures

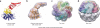

References
-
- Dobson CM. Principles of protein folding, misfolding and aggregation. Semin. Cell Dev. Biol. 2004;15:3–16. - PubMed
-
- Gregersen N, Bross P, Jorgensen MM, Corydon TJ, Andresen BS. Defective folding and rapid degradation of mutant proteins is a common disease mechanism in genetic disorders. J. Inherit. Metab. Dis. 2000;23:441–447. - PubMed
-
- Frydman J. Folding of newly translated proteins in vivo: the role of molecular chaperones. Annu. Rev. Biochem. 2001;70:603–647. - PubMed
-
- Hartl FU, Hayer-Hartl M. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science. 2002;295:1852–1858. - PubMed
-
- Sigler PB, et al. Structure and function in GroEL-mediated protein folding. Annu. Rev. Biochem. 1998;67:581–608. - PubMed

