Overlapping protein-encoding genes in Pseudomonas fluorescens Pf0-1
- PMID: 18551168
- PMCID: PMC2396522
- DOI: 10.1371/journal.pgen.1000094
Overlapping protein-encoding genes in Pseudomonas fluorescens Pf0-1
Abstract
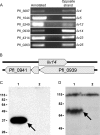
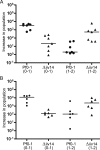
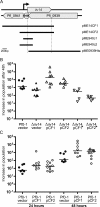
The annotated genome sequences of prokaryotes seldom include overlapping genes encoded opposite each other by the same stretch of DNA. However, antisense transcription is becoming recognized as a widespread phenomenon in eukaryotes, and examples have been linked to important biological processes. Pseudomonas fluorescens inhabits aquatic and terrestrial environments, and can be regarded as an environmental generalist. The genetic basis for this ecological success is not well understood. In a previous search for soil-induced genes in P. fluorescens Pf0-1, ten antisense genes were discovered. These were termed 'cryptic' genes, as they had escaped detection by gene-hunting algorithms, and lacked easily recognizable promoters. In this communication, we designate such genes as 'non-predicted' or 'hidden'. Using reverse transcription PCR, we show that at each of six non-predicted gene loci chosen for study, transcription occurs from both 'sense' and 'antisense' DNA strands. Further, at least one of these hidden antisense genes, iiv14, encodes a protein, as does the sense transcript, both identified by poly-histidine tags on the C-terminus of the proteins. Mutational and complementation studies showed that this novel antisense gene was important for efficient colonization of soil, and multiple copies in the wildtype host improved the speed of soil colonization. Introduction of a stop codon early in the gene eliminated complementation, further implicating the protein in colonization of soil. We therefore designate iiv14 "cosA". These data suggest that, as is the case with eukaryotes, some bacterial genomes are more densely coded than currently recognized.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

References
-
- Rainey PB. Adaptation of Pseudomonas fluorescens to the plant rhizosphere. Environ Microbiol. 1999;1:243–257. - PubMed
-
- Brown DG, Allen C. Ralstonia solanacearum genes induced during growth in tomato: an inside view of bacterial wilt. Mol Microbiol. 2004;53:1641–1660. - PubMed
-
- Gal M, Preston GM, Massey RC, Spiers AJ, Rainey PB. Genes encoding a cellulosic polymer contribute toward the ecological success of Pseudomonas fluorescens SBW25 on plant surfaces. Mol Ecol. 2003;12:3109–3121. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

