Ameobal pathogen mimivirus infects macrophages through phagocytosis
- PMID: 18551172
- PMCID: PMC2398789
- DOI: 10.1371/journal.ppat.1000087
Ameobal pathogen mimivirus infects macrophages through phagocytosis
Abstract
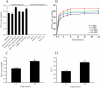
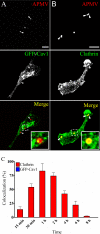
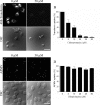
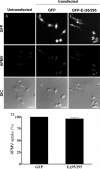
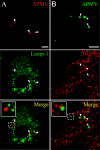
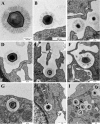
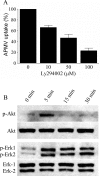
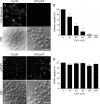
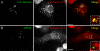
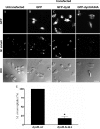
Mimivirus, or Acanthamoeba polyphaga mimivirus (APMV), a giant double-stranded DNA virus that grows in amoeba, was identified for the first time in 2003. Entry by phagocytosis within amoeba has been suggested but not demonstrated. We demonstrate here that APMV was internalized by macrophages but not by non-phagocytic cells, leading to productive APMV replication. Clathrin- and caveolin-mediated endocytosis pathways, as well as degradative endosome-mediated endocytosis, were not used by APMV to invade macrophages. Ultrastructural analysis showed that protrusions were formed around the entering virus, suggesting that macropinocytosis or phagocytosis was involved in APMV entry. Reorganization of the actin cytoskeleton and activation of phosphatidylinositol 3-kinases were required for APMV entry. Blocking macropinocytosis and the lack of APMV colocalization with rabankyrin-5 showed that macropinocytosis was not involved in viral entry. Overexpression of a dominant-negative form of dynamin-II, a regulator of phagocytosis, inhibited APMV entry. Altogether, our data demonstrated that APMV enters macrophages through phagocytosis, a new pathway for virus entry in cells. This reinforces the paradigm that intra-amoebal pathogens have the potential to infect macrophages.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Similar articles
-
Mimivirus: leading the way in the discovery of giant viruses of amoebae.Nat Rev Microbiol. 2017 Apr;15(4):243-254. doi: 10.1038/nrmicro.2016.197. Epub 2017 Feb 27. Nat Rev Microbiol. 2017. PMID: 28239153 Free PMC article. Review.
-
Sputnik virophage disrupts the transcriptional regulation of its host giant virus.J Virol. 2025 Apr 15;99(4):e0019225. doi: 10.1128/jvi.00192-25. Epub 2025 Mar 12. J Virol. 2025. PMID: 40071923 Free PMC article.
-
The virophage as a unique parasite of the giant mimivirus.Nature. 2008 Sep 4;455(7209):100-4. doi: 10.1038/nature07218. Nature. 2008. PMID: 18690211
-
Amoebae, Giant Viruses, and Virophages Make Up a Complex, Multilayered Threesome.Front Cell Infect Microbiol. 2018 Jan 11;7:527. doi: 10.3389/fcimb.2017.00527. eCollection 2017. Front Cell Infect Microbiol. 2018. PMID: 29376032 Free PMC article. Review.
-
Amoebas as mimivirus bunkers: increased resistance to UV light, heat and chemical biocides when viruses are carried by amoeba hosts.Arch Virol. 2014 May;159(5):1039-43. doi: 10.1007/s00705-013-1924-z. Epub 2013 Nov 26. Arch Virol. 2014. PMID: 24271007
Cited by
-
Zinc Finger Protein BCL11A Contributes to the Abortive Infection of Hirame novirhabdovirus (HIRRV) in B Lymphocytes of Flounder (Paralichthys olivaceus).J Virol. 2022 Dec 21;96(24):e0147022. doi: 10.1128/jvi.01470-22. Epub 2022 Nov 30. J Virol. 2022. PMID: 36448803 Free PMC article.
-
A Brief History of Giant Viruses' Studies in Brazilian Biomes.Viruses. 2022 Jan 19;14(2):191. doi: 10.3390/v14020191. Viruses. 2022. PMID: 35215784 Free PMC article. Review.
-
Structural and Proteomic Characterization of the Initiation of Giant Virus Infection.Cell. 2020 May 28;181(5):1046-1061.e6. doi: 10.1016/j.cell.2020.04.032. Epub 2020 May 8. Cell. 2020. PMID: 32392465 Free PMC article.
-
Zamilon, a novel virophage with Mimiviridae host specificity.PLoS One. 2014 Apr 18;9(4):e94923. doi: 10.1371/journal.pone.0094923. eCollection 2014. PLoS One. 2014. PMID: 24747414 Free PMC article.
-
Promoting AMPK/SR-A1-mediated clearance of HMGB1 attenuates chemotherapy-induced peripheral neuropathy.Cell Commun Signal. 2023 May 4;21(1):99. doi: 10.1186/s12964-023-01100-9. Cell Commun Signal. 2023. PMID: 37143083 Free PMC article.
References
-
- La Scola B, Audic S, Robert C, Jungang L, de Lamballerie X, et al. A giant virus in amoebae. Science. 2003;299:2033. - PubMed
-
- Raoult D, La Scola B, Birtles R. The discovery and characterization of Mimivirus, the largest known virus and putative pneumonia agent. Clin Infect Dis. 2007;45:95–102. - PubMed
-
- Khan M, La Scola B, Lepidi H, Raoult D. Pneumonia in mice inoculated experimentally with Acanthamoeba polyphaga mimivirus. Microb Pathog. 2007;42:56–61. - PubMed
-
- Xiao C, Chipman PR, Battisti AJ, Bowman VD, Renesto P, et al. Cryo-electron microscopy of the giant Mimivirus. J Mol Biol. 2005;353:493–496. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

