Enhancement of the alcoholytic activity of alpha-amylase AmyA from Thermotoga maritima MSB8 (DSM 3109) by site-directed mutagenesis
- PMID: 18552192
- PMCID: PMC2519283
- DOI: 10.1128/AEM.00121-08
Enhancement of the alcoholytic activity of alpha-amylase AmyA from Thermotoga maritima MSB8 (DSM 3109) by site-directed mutagenesis
Abstract
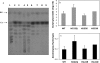
AmyA, an alpha-amylase from the hyperthermophilic bacterium Thermotoga maritima, is able to hydrolyze internal alpha-1,4-glycosidic bonds in various alpha-glucans at 85 degrees C as the optimal temperature. Like other glycoside hydrolases, AmyA also catalyzes transglycosylation reactions, particularly when oligosaccharides are used as substrates. It was found that when methanol or butanol was used as the nucleophile instead of water, AmyA was able to catalyze alcoholysis reactions. This capability has been evaluated in the past for some alpha-amylases, with the finding that only the saccharifying fungal amylases from Aspergillus niger and from Aspergillus oryzae present measurable alcoholysis activity (R. I. Santamaria, G. Del Rio, G. Saab, M. E. Rodriguez, X. Soberon, and A. Lopez, FEBS Lett. 452:346-350, 1999). In the present work, we found that AmyA generates larger quantities of alkyl glycosides than any amylase reported so far. In order to increase the alcoholytic activity observed in AmyA, several residues were identified and mutated based on previous analogous positions in amylases, defining the polarity and geometry of the active site. Replacement of residue His222 by glutamine generated an increase in the alkyl glucoside yield as a consequence of a higher alcoholysis/hydrolysis ratio. The same change in specificity was observed for the mutants H222E and H222D, but instability of these mutants toward alcohols decreased the yield of alkyl glucoside.
Figures
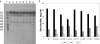

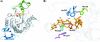



References
-
- Aghajari, N., M. Roth, and R. Haser. 2002. Crystallographic evidence of a transglycosylation reaction: ternary complexes of a psychrophilic alpha-amylase. Biochemistry 41:4273-4280. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

