Conditional mouse osteosarcoma, dependent on p53 loss and potentiated by loss of Rb, mimics the human disease
- PMID: 18559481
- PMCID: PMC2428063
- DOI: 10.1101/gad.1656808
Conditional mouse osteosarcoma, dependent on p53 loss and potentiated by loss of Rb, mimics the human disease
Abstract
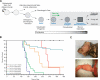
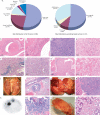
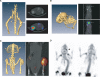
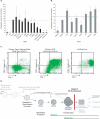
Osteosarcoma is the most common primary malignant tumor of bone. Analysis of familial cancer syndromes and sporadic cases has strongly implicated both p53 and pRb in its pathogenesis; however, the relative contribution of these mutations to the initiation of osteosarcoma is unclear. We describe here the generation and characterization of a genetically engineered mouse model in which all animals develop short latency malignant osteosarcoma. The genetically engineered mouse model is based on osteoblast-restricted deletion of p53 and pRb. Osteosarcoma development is dependent on loss of p53 and potentiated by loss of pRb, revealing a dominance of p53 mutation in the development of osteosarcoma. The model reproduces many of the defining features of human osteosarcoma including cytogenetic complexity and comparable gene expression signatures, histology, and metastatic behavior. Using a novel in silico methodology termed cytogenetic region enrichment analysis, we demonstrate high conservation of gene expression changes between murine osteosarcoma and known cytogentically rearranged loci from human osteosarcoma. Due to the strong similarity between murine osteosarcoma and human osteosarcoma in this model, this should provide a valuable platform for addressing the molecular genetics of osteosarcoma and for developing novel therapeutic strategies.
Figures
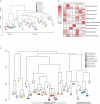

References
-
- Aubin J.E. Regulation of osteoblast formation and function. Rev. Endocr. Metab. Disord. 2001;2:81–94. - PubMed
-
- Becher O.J., Holland E.C. Genetically engineered models have advantages over xenografts for preclinical studies. Cancer Res. 2006;66:3355–3358. - PubMed
-
- Brenner W., Vernon C., Muzi M., Mankoff D.A., Link J.M., Conrad E.U., Eary J.F. Comparison of different quantitative approaches to 18F-fluoride PET scans. J. Nucl. Med. 2004;45:1493–1500. - PubMed
-
- Bridge J.A., Bhatia P.S., Anderson J.R., Neff J.R. Biologic and clinical significance of cytogenetic and molecular cytogenetic abnormalities in benign and malignant cartilaginous lesions. Cancer Genet. Cytogenet. 1993;69:79–90. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous
