Mediators of reprogramming: transcription factors and transitions through mitosis
- PMID: 18568039
- PMCID: PMC7250051
- DOI: 10.1038/nrm2439
Mediators of reprogramming: transcription factors and transitions through mitosis
Abstract
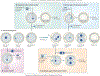
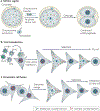
It is thought that most cell types of the human body share the same genetic information as that contained in the zygote from which they originate. Consistent with this view, animal cloning studies demonstrated that the intact genome of a differentiated cell can be reprogrammed to support the development of an entire organism and allow the production of pluripotent stem cells. Recent progress in reprogramming research now points to an important role for transcription factors in the establishment and the maintenance of cellular phenotypes, and to cell division as a mediator of transitions between different states of gene expression.
Figures



Similar articles
-
Asymmetric reprogramming capacity of parental pronuclei in mouse zygotes.Cell Rep. 2014 Mar 27;6(6):1008-1016. doi: 10.1016/j.celrep.2014.02.018. Epub 2014 Mar 13. Cell Rep. 2014. PMID: 24630997
-
Reprogramming of somatic cell identity.Cold Spring Harb Symp Quant Biol. 2008;73:147-55. doi: 10.1101/sqb.2008.73.025. Epub 2008 Nov 6. Cold Spring Harb Symp Quant Biol. 2008. PMID: 19022760 Review.
-
Nuclear reprogramming and pluripotency.Nature. 2006 Jun 29;441(7097):1061-7. doi: 10.1038/nature04955. Nature. 2006. PMID: 16810240 Review.
-
How to improve the success rate of mouse cloning technology.J Reprod Dev. 2010 Feb;56(1):20-30. doi: 10.1262/jrd.09-221a. J Reprod Dev. 2010. PMID: 20203432 Review.
-
Variable reprogramming of the pluripotent stem cell marker Oct4 in mouse clones: distinct developmental potentials in different culture environments.Stem Cells. 2005 Sep;23(8):1089-104. doi: 10.1634/stemcells.2004-0352. Epub 2005 Jun 14. Stem Cells. 2005. PMID: 15955835
Cited by
-
Cell division requires RNA eviction from condensing chromosomes.J Cell Biol. 2020 Nov 2;219(11):e201910148. doi: 10.1083/jcb.201910148. J Cell Biol. 2020. PMID: 33053167 Free PMC article.
-
Asymmetric Regulation of Peripheral Genes by Two Transcriptional Regulatory Networks.PLoS One. 2016 Aug 2;11(8):e0160459. doi: 10.1371/journal.pone.0160459. eCollection 2016. PLoS One. 2016. PMID: 27483142 Free PMC article.
-
Reprogramming: Remodelling for pluripotency.Nat Rev Mol Cell Biol. 2010 Aug;11(8):540-1. doi: 10.1038/nrm2942. Epub 2010 Jul 14. Nat Rev Mol Cell Biol. 2010. PMID: 20628409 No abstract available.
-
Cloning by SCNT: Integrating Technical and Biology-Driven Advances.Methods Mol Biol. 2023;2647:1-35. doi: 10.1007/978-1-0716-3064-8_1. Methods Mol Biol. 2023. PMID: 37041327
-
A theoretical model of neural maturation in the developing chick spinal cord.PLoS One. 2020 Dec 18;15(12):e0244219. doi: 10.1371/journal.pone.0244219. eCollection 2020. PLoS One. 2020. PMID: 33338079 Free PMC article.
References
-
-
Gurdon JB, Elsdale TR & Fischberg M. Sexually mature individuals of Xenopus laevis from the transplantation of single somatic nuclei. Nature 182, 64–65 (1958).
Demonstrates that frogs can be cloned from somatic cells.
-
-
-
Wilmut I, Schnieke AE, McWhir J, Kinnotd AJ & Campbell KH Viable offspring derived from fetal and adult mammalian cells. Nature 385, 810–813 (1997).
Demonstrates the reversibility of mammalian differentiation.
-
-
- Eggan K. et al. Mice cloned from olfactory sensory neurons. Nature 428, 44–49 (2004). - PubMed
-
-
Hochedlingoteer K. & Jaenisch R. Monoclonal mice generated by nuclear transfer from mature B and T donor cells. Nature 415, 1035–1038 (2002).
Demonstrates that the genome of even the most differentiated cells can support development after nuclear transfer.
-
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

