Regulation of ABCG2 expression at the 3' untranslated region of its mRNA through modulation of transcript stability and protein translation by a putative microRNA in the S1 colon cancer cell line
- PMID: 18573883
- PMCID: PMC2519722
- DOI: 10.1128/MCB.00331-08
Regulation of ABCG2 expression at the 3' untranslated region of its mRNA through modulation of transcript stability and protein translation by a putative microRNA in the S1 colon cancer cell line
Abstract
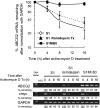
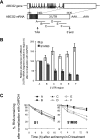
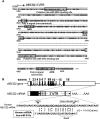
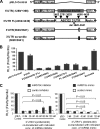
ABCG2 is recognized as an important efflux transporter in clinical pharmacology and is potentially important in resistance to chemotherapeutic drugs. To identify epigenetic mechanisms regulating ABCG2 mRNA expression at its 3' untranslated region (3'UTR), we performed 3' rapid amplification of cDNA ends with the S1 parental colon cancer cell line and its drug-resistant ABCG2-overexpressing counterpart. We found that the 3'UTR is >1,500 bp longer in parental cells and, using the miRBase TARGETs database, identified a putative microRNA (miRNA) binding site, distinct from the recently reported hsa-miR520h site, in the portion of the 3'UTR missing from ABCG2 mRNA in the resistant cells. We hypothesized that the binding of a putative miRNA at the 3'UTR of ABCG2 suppresses the expression of ABCG2. In resistant S1MI80 cells, the miRNA cannot bind to ABCG2 mRNA because of the shorter 3'UTR, and thus, mRNA degradation and/or repression on protein translation is relieved, contributing to overexpression of ABCG2. This hypothesis was rigorously tested by reporter gene assays, mutational analysis at the miRNA binding sites, and forced expression of miRNA inhibitors or mimics. The removal of this epigenetic regulation by miRNA could be involved in the overexpression of ABCG2 in drug-resistant cancer cells.
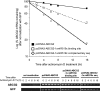
Figures




Similar articles
-
Escape from hsa-miR-519c enables drug-resistant cells to maintain high expression of ABCG2.Mol Cancer Ther. 2009 Oct;8(10):2959-68. doi: 10.1158/1535-7163.MCT-09-0292. Mol Cancer Ther. 2009. PMID: 19825807 Free PMC article.
-
The C421A (Q141K) polymorphism enhances the 3'-untranslated region (3'-UTR)-dependent regulation of ATP-binding cassette transporter ABCG2.Biochem Pharmacol. 2016 Mar 15;104:139-47. doi: 10.1016/j.bcp.2016.02.011. Epub 2016 Feb 21. Biochem Pharmacol. 2016. PMID: 26903388
-
Breast cancer resistance protein BCRP/ABCG2 regulatory microRNAs (hsa-miR-328, -519c and -520h) and their differential expression in stem-like ABCG2+ cancer cells.Biochem Pharmacol. 2011 Mar 15;81(6):783-92. doi: 10.1016/j.bcp.2010.12.018. Epub 2011 Jan 8. Biochem Pharmacol. 2011. PMID: 21219875 Free PMC article.
-
MicroRNA: a prognostic biomarker and a possible druggable target for circumventing multidrug resistance in cancer chemotherapy.J Biomed Sci. 2013 Dec 20;20(1):99. doi: 10.1186/1423-0127-20-99. J Biomed Sci. 2013. PMID: 24358977 Free PMC article. Review.
-
Regulation and dysregulation of 3'UTR-mediated translational control.Curr Opin Genet Dev. 2013 Feb;23(1):29-34. doi: 10.1016/j.gde.2012.12.004. Epub 2013 Jan 9. Curr Opin Genet Dev. 2013. PMID: 23312843 Free PMC article. Review.
Cited by
-
MiRNA-149 modulates chemosensitivity of ovarian cancer A2780 cells to paclitaxel by targeting MyD88.J Ovarian Res. 2015 Jul 30;8:48. doi: 10.1186/s13048-015-0178-7. J Ovarian Res. 2015. PMID: 26223974 Free PMC article.
-
Transporters in the Mammary Gland-Contribution to Presence of Nutrients and Drugs into Milk.Nutrients. 2019 Oct 5;11(10):2372. doi: 10.3390/nu11102372. Nutrients. 2019. PMID: 31590349 Free PMC article. Review.
-
MicroRNAs and their relevance to ABC transporters.Br J Clin Pharmacol. 2014 Apr;77(4):587-96. doi: 10.1111/bcp.12251. Br J Clin Pharmacol. 2014. PMID: 24645868 Free PMC article. Review.
-
[Advances in tumor chemo-resistance regulated by MicroRNA].Zhongguo Fei Ai Za Zhi. 2014 Oct 20;17(10):741-9. doi: 10.3779/j.issn.1009-3419.2014.10.06. Zhongguo Fei Ai Za Zhi. 2014. PMID: 25342041 Free PMC article. Review. Chinese.
-
MicroRNA 130b enhances drug resistance in human ovarian cancer cells.Tumour Biol. 2014 Dec;35(12):12151-6. doi: 10.1007/s13277-014-2520-x. Epub 2014 Aug 26. Tumour Biol. 2014. PMID: 25155039
References
-
- Allikmets, R., L. M. Schriml, A. Hutchinson, V. Romano-Spica, and M. Dean. 1998. A human placenta-specific ATP-binding cassette gene (ABCP) on chromosome 4q22 that is involved in multidrug resistance. Cancer Res. 585337-5339. - PubMed
-
- Ambros, V. 2004. The functions of animal microRNAs. Nature 431350-355. - PubMed
-
- Bandres, E., E. Cubedo, X. Agirre, R. Malumbres, R. Zarate, N. Ramirez, A. Abajo, A. Navarro, I. Moreno, M. Monzo, and J. Garcia-Foncillas. 2006. Identification by real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol. Cancer 529. - PMC - PubMed
-
- Basler, C. F., and M. S. Horwitz. 1996. Subgroup B adenovirus type 35 early region 3 mRNAs differ from those of the subgroup C adenoviruses. Virology 215165-177. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
