A flexible template boundary element in the RNA subunit of fission yeast telomerase
- PMID: 18574244
- PMCID: PMC2527122
- DOI: 10.1074/jbc.M802043200
A flexible template boundary element in the RNA subunit of fission yeast telomerase
Abstract
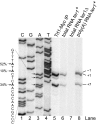
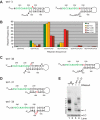
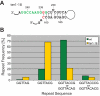
Telomerase adds telomeric repeat sequences to chromosome ends using a short region of its RNA subunit as a template. Telomerase RNA subunits are phylogenetically highly divergent, and different strategies have evolved to demarcate the boundary of the template region. The recent identification of the gene encoding telomerase RNA in the fission yeast Schizosaccharomyces pombe (ter1+) has opened the door for structure-function analyses in a model that shares many features with the telomere maintenance machinery of higher eukaryotes. Here we describe a structural element in TER1 that defines the 5' boundary of the template. Disruption of a predicted long range base pairing interaction between template-adjacent nucleotides and a sequence further upstream resulted in reverse transcription beyond the template region and caused telomere shortening. Normal telomere length was restored by combining complementary nucleotide substitutions in both elements, showing that base pairing, not a specific sequence, limits reverse transcription beyond the template. The template boundary described here resembles that of budding yeasts and some mammalian telomerases. However, unlike any previously characterized boundary element, part of the paired region overlaps with the template itself, thus necessitating disruption of these interactions during most reverse transcription cycles. We show that changes in the paired region directly affect the length of individual telomeric repeat units. Our data further illustrate that marginal alignment of the telomeric 3' end with RNA sequences downstream of the template is responsible for primer slippage, causing incorporation of strings of guanosines at the start of a subset of repeats.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

