The role of retinoblastoma-associated proteins 46 and 48 in estrogen receptor alpha mediated gene expression
- PMID: 18577416
- PMCID: PMC2642675
- DOI: 10.1016/j.mce.2008.05.016
The role of retinoblastoma-associated proteins 46 and 48 in estrogen receptor alpha mediated gene expression
Abstract
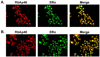
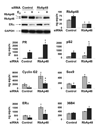
The differential recruitment of coregulatory proteins to the DNA-bound estrogen receptor alpha (ERalpha) plays a critical role in mediating estrogen-responsive gene expression. We previously isolated and identified retinoblastoma-associated proteins 46 (RbAp46) and 48 (RbAp48), which are associated with chromatin remodeling, histone deacetylation, and transcription repression, as proteins associated with the DNA-bound ERalpha. We now demonstrate that RbAp46 and RbAp48 interact with ERalphain vitro and in vivo, associate with ERalpha at endogenous, estrogen-responsive genes, and alter expression of endogenous, ERalpha-activated and -repressed genes in MCF-7 breast cancer cells. Our findings reveal that RbAp48 limits expression of estrogen-responsive genes and that RbAp46 modulates estrogen responsiveness in a gene-specific manner. The ability of RbAp46 and RbAp48 to interact with ERalpha and influence its activity reveals yet another role for these multifunctional proteins in regulating gene expression.
Figures




Similar articles
-
Dual retinoblastoma-binding proteins with properties related to a negative regulator of ras in yeast.J Biol Chem. 1995 Oct 27;270(43):25507-13. doi: 10.1074/jbc.270.43.25507. J Biol Chem. 1995. PMID: 7503932
-
The silencing mediator of retinoic acid and thyroid hormone receptor (SMRT) corepressor is required for full estrogen receptor alpha transcriptional activity.Mol Cell Biol. 2007 Sep;27(17):5933-48. doi: 10.1128/MCB.00237-07. Epub 2007 Jun 25. Mol Cell Biol. 2007. PMID: 17591692 Free PMC article.
-
The histone deacetylase HDAC3 targets RbAp48 to the retinoblastoma protein.Nucleic Acids Res. 2001 Aug 1;29(15):3131-6. doi: 10.1093/nar/29.15.3131. Nucleic Acids Res. 2001. PMID: 11470869 Free PMC article.
-
Amplification of WHSC1L1 regulates expression and estrogen-independent activation of ERα in SUM-44 breast cancer cells and is associated with ERα over-expression in breast cancer.Mol Oncol. 2016 Jun;10(6):850-65. doi: 10.1016/j.molonc.2016.02.003. Epub 2016 Feb 27. Mol Oncol. 2016. PMID: 27005559 Free PMC article.
-
RbAp46 inhibits estrogen-stimulated progression of neoplastigenic breast epithelial cells.Anticancer Res. 2007 Sep-Oct;27(5A):3205-9. Anticancer Res. 2007. PMID: 17970062
Cited by
-
The multifaceted histone chaperone RbAp46/48 in Plasmodium falciparum: structural insights, production, and characterization.Parasitol Res. 2020 Jun;119(6):1753-1765. doi: 10.1007/s00436-020-06669-5. Epub 2020 May 4. Parasitol Res. 2020. PMID: 32363442
-
Unveiling the molecular structure and role of RBBP4/7: implications for epigenetic regulation and cancer research.Front Mol Biosci. 2023 Nov 13;10:1276612. doi: 10.3389/fmolb.2023.1276612. eCollection 2023. Front Mol Biosci. 2023. PMID: 38028543 Free PMC article. Review.
-
Retinoblastoma treatment: impact of the glycolytic inhibitor 2-deoxy-d-glucose on molecular genomics expression in LH(BETA)T(AG) retinal tumors.Clin Ophthalmol. 2012;6:817-30. doi: 10.2147/OPTH.S29688. Epub 2012 May 29. Clin Ophthalmol. 2012. PMID: 22701083 Free PMC article.
-
Apurinic/apyrimidinic endonuclease 1 alters estrogen receptor activity and estrogen-responsive gene expression.Mol Endocrinol. 2009 Sep;23(9):1346-59. doi: 10.1210/me.2009-0093. Epub 2009 May 21. Mol Endocrinol. 2009. PMID: 19460860 Free PMC article.
-
The role of maintenance proteins in the preservation of epithelial cell identity during mammary gland remodeling and breast cancer initiation.Chin J Cancer. 2014 Feb;33(2):51-67. doi: 10.5732/cjc.013.10040. Epub 2013 Jul 12. Chin J Cancer. 2014. PMID: 23845141 Free PMC article. Review.
References
-
- Ahringer J. NuRD and SIN3 histone deacetylase complexes in development. Trends Genet. 2000;16:351–356. - PubMed
-
- Bowen NJ, Fujita N, Kajita M, Wade PA. Mi-2/NuRD: multiple complexes for many purposes. Biochim Biophys Acta. 2004;1677:52–57. - PubMed
-
- Carroll J, Meyer C, Song J, Li W, Geistlinger T, Eeckhoute J, Brodshy A, Keeton E, Fertuck K, Hall G, Wang Q, Bekiranov S, Sementchenko V, Fox E, Silver P, Gingeras T, Liu X, Brown M. Genome-wide analysis of estrogen receptor binding sites. Nat Genetics. 2006;38:1289–1297. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

