Mutations of glucocorticoid receptor differentially affect AF2 domain activity in a steroid-selective manner to alter the potency and efficacy of gene induction and repression
- PMID: 18578507
- PMCID: PMC2678802
- DOI: 10.1021/bi800472w
Mutations of glucocorticoid receptor differentially affect AF2 domain activity in a steroid-selective manner to alter the potency and efficacy of gene induction and repression
Abstract
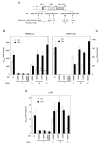
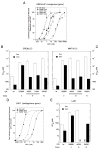
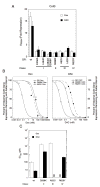
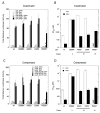
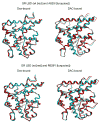
The transcriptional activity of steroid hormones is intimately associated with their structure. Deacylcortivazol (DAC) contains several features that were predicted to make it an inactive glucocorticoid. Nevertheless, gene induction and repression by complexes of glucocorticoid receptor (GR) with DAC occur with potency (lower EC 50) greater than and efficacy (maximal activity, or A max) equal to those of the very active and smaller synthetic glucocorticoid dexamethasone (Dex). Guided by a recent X-ray structure of DAC bound to the GR ligand binding domain (LBD), we now report that several point mutants in the LBD have little effect on the binding of either agonist steroid. However, these same mutations dramatically alter the A max and/or EC 50 of exogenous and endogenous genes in a manner that depends on steroid structure. In some cases, Dex is no longer a full agonist. These properties appear to result from a preferential inactivation of the AF2 activation domain in the GR LBD of Dex-bound, but not DAC-bound, receptors. The Dex-bound receptors display normal binding to, but a greatly reduced response to, the coactivator TIF2, thus indicating a defect in the transmission efficiency of GR-steroid complex information to the coactivator TIF2. In addition, all GR mutants that are active in gene induction with either Dex or DAC have greatly reduced activity in gene repression. This contrasts with the reports of GR mutations preferentially suppressing GR-mediated induction. The properties of these GR mutants in gene induction support the hypothesis that the A max and EC 50 of GR-controlled gene expression can be independently modified, indicate that the receptor can be modified to favor activity with a specific agonist steroid, and suggest that new ligands with suitable substituents may be able to affect the same LBD conformational changes and thereby broaden the therapeutic applications of glucocorticoid steroids.
Figures
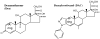
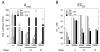
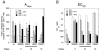
Similar articles
-
Ligand binding domain mutations of the glucocorticoid receptor selectively modify the effects with, but not binding of, cofactors.Biochemistry. 2011 Jan 25;50(3):356-66. doi: 10.1021/bi101792d. Epub 2010 Dec 30. Biochemistry. 2011. PMID: 21142156 Free PMC article.
-
Binding of the N-terminal region of coactivator TIF2 to the intrinsically disordered AF1 domain of the glucocorticoid receptor is accompanied by conformational reorganizations.J Biol Chem. 2012 Dec 28;287(53):44546-60. doi: 10.1074/jbc.M112.411330. Epub 2012 Nov 6. J Biol Chem. 2012. PMID: 23132854 Free PMC article.
-
Glucocorticoid receptor ligand binding domain is sufficient for the modulation of glucocorticoid induction properties by homologous receptors, coactivator transcription intermediary factor 2, and Ubc9.Mol Endocrinol. 2005 Feb;19(2):290-311. doi: 10.1210/me.2004-0134. Epub 2004 Nov 11. Mol Endocrinol. 2005. PMID: 15539428
-
Glucocorticoid receptor cofactors as therapeutic targets.Curr Opin Pharmacol. 2010 Dec;10(6):613-9. doi: 10.1016/j.coph.2010.08.001. Epub 2010 Aug 26. Curr Opin Pharmacol. 2010. PMID: 20801081 Free PMC article. Review.
-
How glucocorticoid receptors modulate the activity of other transcription factors: a scope beyond tethering.Mol Cell Endocrinol. 2013 Nov 5;380(1-2):41-54. doi: 10.1016/j.mce.2012.12.014. Epub 2012 Dec 23. Mol Cell Endocrinol. 2013. PMID: 23267834 Review.
Cited by
-
The glucocorticoid receptor dimer interface allosterically transmits sequence-specific DNA signals.Nat Struct Mol Biol. 2013 Jul;20(7):876-83. doi: 10.1038/nsmb.2595. Epub 2013 Jun 2. Nat Struct Mol Biol. 2013. PMID: 23728292 Free PMC article.
-
The road less traveled: new views of steroid receptor action from the path of dose-response curves.Mol Cell Endocrinol. 2012 Jan 30;348(2):373-82. doi: 10.1016/j.mce.2011.05.030. Epub 2011 Jun 1. Mol Cell Endocrinol. 2012. PMID: 21664235 Free PMC article. Review.
-
STAMP alters the growth of transformed and ovarian cancer cells.BMC Cancer. 2010 Apr 7;10:128. doi: 10.1186/1471-2407-10-128. BMC Cancer. 2010. PMID: 20374646 Free PMC article.
-
Separate regions of glucocorticoid receptor, coactivator TIF2, and comodulator STAMP modify different parameters of glucocorticoid-mediated gene induction.Mol Cell Endocrinol. 2012 May 15;355(1):121-34. doi: 10.1016/j.mce.2012.02.001. Epub 2012 Feb 11. Mol Cell Endocrinol. 2012. PMID: 22342989 Free PMC article.
-
RSUME enhances glucocorticoid receptor SUMOylation and transcriptional activity.Mol Cell Biol. 2013 Jun;33(11):2116-27. doi: 10.1128/MCB.01470-12. Epub 2013 Mar 18. Mol Cell Biol. 2013. PMID: 23508108 Free PMC article.
References
-
- Williams SP, Sigler PB. Atomic structure of progesterone complexed with its receptor. Nature. 1998;393:392–396. - PubMed
-
- Bledsoe RK, Montana VG, Stanley TB, Delves CJ, Apolito CJ, McKee DD, Consler TG, Parks DJ, Stewart EL, Willson TM, Lambert MH, Moore JT, Pearce KH, Xu HE. Crystal structure of the glucocorticoid receptor ligand binding domain reveals a novel mode of receptor dimerization and coactivator recognition. Cell. 2002;110:93–105. - PubMed
-
- Brzozowski AM, Pike ACW, Dauter Z, Hubbard RE, Bonn T, Engstrom O, Ohman L, Greene GL, Gustafsson J-A, Carlquist M. Molecular basis of agonism and antagonism in the oestrogen receptor. Nature. 1997;389:753–758. - PubMed
-
- Shiau AK, Barstad D, Loria PM, Cheng L, Kushner PJ, Agard DA, Greene GL. The structural basis of estrogen receptor/coactivator recognition and the antagonism of this interaction by tamoxifen. Cell. 1998;95:927–937. - PubMed
-
- Kauppi B, Jakob C, Farnegardh M, Yang J, Ahola H, Alarcon M, Calles K, Engstrom O, Harlan J, Muchmore S, Ramqvist AK, Thorell S, Ohman L, Greer J, Gustafsson JA, Carlstedt-Duke J, Carlquist M. The three-dimensional structures of antagonistic and agonistic forms of the glucocorticoid receptor ligand-binding domain: RU-486 induces a transconformation that leads to active antagonism. J Biol Chem. 2003;278:22748–22754. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

