Combinatorial influence of environmental parameters on transcription factor activity
- PMID: 18586711
- PMCID: PMC2718633
- DOI: 10.1093/bioinformatics/btn155
Combinatorial influence of environmental parameters on transcription factor activity
Abstract
Motivation: Cells receive a wide variety of environmental signals, which are often processed combinatorially to generate specific genetic responses. Changes in transcript levels, as observed across different environmental conditions, can, to a large extent, be attributed to changes in the activity of transcription factors (TFs). However, in unraveling these transcription regulation networks, the actual environmental signals are often not incorporated into the model, simply because they have not been measured. The unquantified heterogeneity of the environmental parameters across microarray experiments frustrates regulatory network inference.
Results: We propose an inference algorithm that models the influence of environmental parameters on gene expression. The approach is based on a yeast microarray compendium of chemostat steady-state experiments. Chemostat cultivation enables the accurate control and measurement of many of the key cultivation parameters, such as nutrient concentrations, growth rate and temperature. The observed transcript levels are explained by inferring the activity of TFs in response to combinations of cultivation parameters. The interplay between activated enhancers and repressors that bind a gene promoter determine the possible up- or downregulation of the gene. The model is translated into a linear integer optimization problem. The resulting regulatory network identifies the combinatorial effects of environmental parameters on TF activity and gene expression.
Availability: The Matlab code is available from the authors upon request.
Supplementary information: Supplementary data are available at Bioinformatics online.
Figures


 , the optimal approximation of the discretized regression coefficients in R. (a) The coefficients in R are derived from a regression analysis, which assesses the influence of cultivation parameters on gene expression by employing these parameters as predictors in the regression model. The discretization procedure maps non-zero regression weights to 1 or −1, depending on their sign. (The schematic representation of R is given for five genes and three cultivation parameters.) (b) The elements of
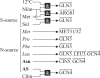
, the optimal approximation of the discretized regression coefficients in R. (a) The coefficients in R are derived from a regression analysis, which assesses the influence of cultivation parameters on gene expression by employing these parameters as predictors in the regression model. The discretization procedure maps non-zero regression weights to 1 or −1, depending on their sign. (The schematic representation of R is given for five genes and three cultivation parameters.) (b) The elements of  are determined by T and M. T is fixed and indicates binary TF binding potential to gene promoters. The elements of M are estimated and indicate the activity of TFs as enhancers or repressors under the different (combinatorial) cultivation parameters. A logic circuit derived from M is graphically depicted above the representation of M. (c) Visualization of the active TFs on the gene promoters of genes g1, g2 and g3 under cultivation parameter A. Enhancers are depicted as red boxes; repressors are depicted as green boxes. (TF γ can bind the promoter of g1, but is not active under A.) The height of a box indicates the enhancer or repressor strength. The strength of a particular enhancer or repressor is the same for all genes. A gene is upregulated when its activator strength, i.e. the sum of the heights of the red boxes, is larger than the repressor strength, which equals the sum of the heights of the green boxes. Downregulation is inferred in the opposite situation. See text for details.
are determined by T and M. T is fixed and indicates binary TF binding potential to gene promoters. The elements of M are estimated and indicate the activity of TFs as enhancers or repressors under the different (combinatorial) cultivation parameters. A logic circuit derived from M is graphically depicted above the representation of M. (c) Visualization of the active TFs on the gene promoters of genes g1, g2 and g3 under cultivation parameter A. Enhancers are depicted as red boxes; repressors are depicted as green boxes. (TF γ can bind the promoter of g1, but is not active under A.) The height of a box indicates the enhancer or repressor strength. The strength of a particular enhancer or repressor is the same for all genes. A gene is upregulated when its activator strength, i.e. the sum of the heights of the red boxes, is larger than the repressor strength, which equals the sum of the heights of the green boxes. Downregulation is inferred in the opposite situation. See text for details.
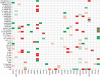
 , indicating the inferred regression coefficients. Upregulation is indicated by red; downregulation is indicated by green. Incorrectly inferred elements are marked with a gray cross. White boxes without a cross are the zero elements of R. These elements are not part of the optimization scheme.
, indicating the inferred regression coefficients. Upregulation is indicated by red; downregulation is indicated by green. Incorrectly inferred elements are marked with a gray cross. White boxes without a cross are the zero elements of R. These elements are not part of the optimization scheme.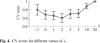
Similar articles
-
Combinatorial effects of environmental parameters on transcriptional regulation in Saccharomyces cerevisiae: a quantitative analysis of a compendium of chemostat-based transcriptome data.BMC Genomics. 2009 Jan 27;10:53. doi: 10.1186/1471-2164-10-53. BMC Genomics. 2009. PMID: 19173729 Free PMC article.
-
Quantitative characterization of the transcriptional regulatory network in the yeast cell cycle.Bioinformatics. 2004 Aug 12;20(12):1914-27. doi: 10.1093/bioinformatics/bth178. Epub 2004 Mar 25. Bioinformatics. 2004. PMID: 15044243
-
Exploiting combinatorial cultivation conditions to infer transcriptional regulation.BMC Genomics. 2007 Jan 22;8:25. doi: 10.1186/1471-2164-8-25. BMC Genomics. 2007. PMID: 17241460 Free PMC article.
-
Chemostat-based micro-array analysis in baker's yeast.Adv Microb Physiol. 2009;54:257-311. doi: 10.1016/S0065-2911(08)00004-0. Adv Microb Physiol. 2009. PMID: 18929070 Review.
-
Inferring network interactions within a cell.Brief Bioinform. 2005 Dec;6(4):380-9. doi: 10.1093/bib/6.4.380. Brief Bioinform. 2005. PMID: 16420736 Review.
Cited by
-
Integrating genome assemblies with MAIA.Bioinformatics. 2010 Sep 15;26(18):i433-9. doi: 10.1093/bioinformatics/btq366. Bioinformatics. 2010. PMID: 20823304 Free PMC article.
-
Stability from structure: metabolic networks are unlike other biological networks.EURASIP J Bioinform Syst Biol. 2009;2009(1):630695. doi: 10.1155/2009/630695. Epub 2009 Feb 1. EURASIP J Bioinform Syst Biol. 2009. PMID: 19197379 Free PMC article.
-
Genome-wide analytical approaches for reverse metabolic engineering of industrially relevant phenotypes in yeast.FEMS Yeast Res. 2012 Mar;12(2):183-96. doi: 10.1111/j.1567-1364.2011.00776.x. Epub 2012 Jan 10. FEMS Yeast Res. 2012. PMID: 22152095 Free PMC article. Review.
References
-
- Balaji S, et al. Comprehensive analysis of combinatorial regulation using the transcriptional regulatory network of yeast. J. Mol. Biol. 2006;360:213–227. - PubMed
-
- Bammler T, et al. Standardizing global gene expression analysis between laboratories and across platforms. Nat. Methods. 2005;2:351–356. - PubMed
-
- Bar-Joseph Z, et al. Computational discovery of gene modules and regulatory networks. Nat. Biotechnol. 2003;21:1337–1342. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

