Predicting protein thermostability changes from sequence upon multiple mutations
- PMID: 18586713
- PMCID: PMC2718644
- DOI: 10.1093/bioinformatics/btn166
Predicting protein thermostability changes from sequence upon multiple mutations
Abstract
Motivation: A basic question in protein science is to which extent mutations affect protein thermostability. This knowledge would be particularly relevant for engineering thermostable enzymes. In several experimental approaches, this issue has been serendipitously addressed. It would be therefore convenient providing a computational method that predicts when a given protein mutant is more thermostable than its corresponding wild-type.
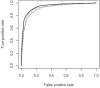
Results: We present a new method based on support vector machines that is able to predict whether a set of mutations (including insertion and deletions) can enhance the thermostability of a given protein sequence. When trained and tested on a redundancy-reduced dataset, our predictor achieves 88% accuracy and a correlation coefficient equal to 0.75. Our predictor also correctly classifies 12 out of 14 experimentally characterized protein mutants with enhanced thermostability. Finally, it correctly detects all the 11 mutated proteins whose increase in stability temperature is >10 degrees C.
Availability: The dataset and the list of protein clusters adopted for the SVM cross-validation are available at the web site http://lipid.biocomp.unibo.it/~ludovica/thermo-meso-MUT.
Figures


References
-
- Annaluru N, et al. Thermostabilization of Pichia stipitis xylitol dehydrogenase by mutation of structural zinc-binding loop. J. Biotechnol. 2007;129:717–722. - PubMed
-
- Bakke M, et al. Thermostabilization of porcine kidney D-amino acid oxidase by a single amino acid substitution. Biotechnol. Bioeng., 2006;93:1023–1027. - PubMed
-
- Bommarius AS, et al. High-throughput screening for enhanced protein stability. Curr. Opin. Biotechnol. 2006;17:606–610. - PubMed
-
- Brouns SJ, et al. Engineering a selectable marker for hyperthermophiles. J. Biol. Chem., 2005;280:11422–11431. - PubMed
-
- Burges CJC. A Tutorial on Support Vector Machines for Pattern Recognition. Boston: Kluwer Academic Publishers; 1998.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

