Single-cell transcription site activation predicts chemotherapy response in human colorectal tumors
- PMID: 18593893
- PMCID: PMC3107669
- DOI: 10.1158/0008-5472.CAN-07-6770
Single-cell transcription site activation predicts chemotherapy response in human colorectal tumors
Abstract
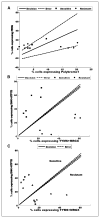
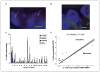
Candidate gene and pathway approaches, and unbiased gene expression profiling, have identified marker signatures predictive of tumor phenotypes, such as drug sensitivity and invasive or metastatic potential. However, application of such information to evaluation of tumors in the clinic is limited by cell heterogeneity in the tumor. We have developed a novel method of fluorescence in situ hybridization (FISH) that can detect transcriptional activation of individual genes at their site in single cells in the interphase nucleus. A major obstacle in the treatment of colorectal cancer is relative insensitivity to the chemotherapeutic agent 5-Fluorouracil (5-FU). Here, we have developed a sensitive approach to predict relative sensitivity of colorectal cancer cells to 5-FU, using FISH with probes targeted to nascent mRNAs to measure the number of individual cells with active transcription sites for a panel of candidate genes. These results reveal that the transcriptional status of four key genes, thymidylate synthase (TYMS), MORF-related gene X (MRGX), Bcl2-antagonist/killer (BAK), and ATPase, Cu(2+) transporting beta polypeptide (ATP7B), can accurately predict response to 5-FU. As proof of principle, we show that this transcriptional profile is predictive of response to 5-FU in a small number of patient colon tumor tissues. This approach provides a novel ability to identify and characterize unique minor cell populations in the tumor that may exhibit relative resistance to chemotherapy.
Conflict of interest statement
L.H. Augenlicht: consultant/advisory board, Pittsburgh Cancer Center, Arizona Cancer Center, and Valley Hospital, New Jersey. R.H. Singer: consultant/advisory board, Aureon Laboratories. The other authors disclosed no potential conflicts of interest.
Figures


References
-
- Longley DB, Harkin DP, Johnston PG. 5-fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer. 2003;3:330–8. - PubMed
-
- Leichman CG, Lenz HJ, Leichman L, et al. Quantitation of intratumoral thymidylate synthase expression predicts for disseminated colorectal cancer response and resistance to protracted-infusion fluorouracil and weekly leucovorin. J Clin Oncol. 1997;15:3223–9. - PubMed
-
- Salonga D, Danenberg KD, Johnson M, et al. Colorectal tumors responding to 5-fluorouracil have low gene expression levels of dihydropyrimidine dehydrogenase, thymidylate synthase, and thymidine phosphorylase. Clin Cancer Res. 2000;6:1322–7. - PubMed
-
- Metzger R, Danenberg K, Leichman CG, et al. High basal level gene expression of thymidine phosphorylase (platelet-derived endothelial cell growth factor) in colorectal tumors is associated with nonresponse to 5-fluorouracil. Clin Cancer Res. 1998;4:2371–6. - PubMed
-
- Elsaleh H, Iacopetta B. Microsatellite instability is a predictive marker for survival benefit from adjuvant chemotherapy in a population-based series of stage III colorectal carcinoma. Clin Colorectal Cancer. 2001;1:104–9. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 CA075503/CA/NCI NIH HHS/United States
- P30CA56036/CA/NCI NIH HHS/United States
- R01CA75503/CA/NCI NIH HHS/United States
- R33CA83208/CA/NCI NIH HHS/United States
- R33 CA083208/CA/NCI NIH HHS/United States
- R01CA107382/CA/NCI NIH HHS/United States
- R01CA70896/CA/NCI NIH HHS/United States
- P30 CA013330/CA/NCI NIH HHS/United States
- R01 CA107382/CA/NCI NIH HHS/United States
- T32GM07288/GM/NIGMS NIH HHS/United States
- R01 CA070896/CA/NCI NIH HHS/United States
- P30CA13330/CA/NCI NIH HHS/United States
- U54CA100926/CA/NCI NIH HHS/United States
- R01CA86072/CA/NCI NIH HHS/United States
- T32 GM007288/GM/NIGMS NIH HHS/United States
- R01 CA086072/CA/NCI NIH HHS/United States
- P30 CA056036/CA/NCI NIH HHS/United States
- U54 CA100926/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Research Materials

