Shared gene expression profiles in developing heart valves and osteoblast progenitor cells
- PMID: 18612084
- PMCID: PMC2536828
- DOI: 10.1152/physiolgenomics.90212.2008
Shared gene expression profiles in developing heart valves and osteoblast progenitor cells
Abstract
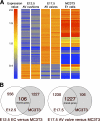
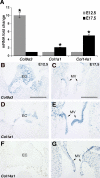
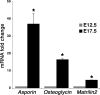
The atrioventricular (AV) valves of the heart develop from undifferentiated mesenchymal endocardial cushions, which later mature into stratified valves with diversified extracellular matrix (ECM). Because the mature valves express genes associated with osteogenesis and exhibit disease-associated calcification, we hypothesized the existence of shared regulatory pathways active in developing AV valves and in bone progenitor cells. To define gene regulatory programs of valvulogenesis relative to osteoblast progenitors, we undertook Affymetrix gene expression profiling analysis of murine embryonic day (E)12.5 AV endocardial cushions compared with E17.5 AV valves (mitral and tricuspid) and with preosteoblast MC3T3-E1 (subclone4) cells. Overall, MC3T3 cells were significantly more similar to E17.5 valves than to E12.5 cushions, supporting the hypothesis that valve maturation involves the expression of many genes also expressed in osteoblasts. Several transcription factors characteristic of mesenchymal and osteoblast precursor cells, including Twist1, are predominant in E12.5 cushion. Valve maturation is characterized by differential regulation of matrix metalloproteinases and their inhibitors as well as complex collagen gene expression. Among the most highly enriched genes during valvulogenesis were members of the small leucine-rich proteoglycan (SLRP) family including Asporin, a known negative regulator of osteoblast differentiation and mineralization. Together, these data support shared gene expression profiles of the developing valves and osteoblast bone precursor cells in normal valve development and homeostasis with potential functions in calcific valve disease.
Figures


Similar articles
-
BMP2 expression in the endocardial lineage is required for AV endocardial cushion maturation and remodeling.Dev Biol. 2017 Oct 1;430(1):113-128. doi: 10.1016/j.ydbio.2017.08.008. Epub 2017 Aug 6. Dev Biol. 2017. PMID: 28790014 Free PMC article.
-
Heart valve development: regulatory networks in development and disease.Circ Res. 2009 Aug 28;105(5):408-21. doi: 10.1161/CIRCRESAHA.109.201566. Circ Res. 2009. PMID: 19713546 Free PMC article. Review.
-
Twist1 promotes heart valve cell proliferation and extracellular matrix gene expression during development in vivo and is expressed in human diseased aortic valves.Dev Biol. 2010 Nov 1;347(1):167-79. doi: 10.1016/j.ydbio.2010.08.021. Epub 2010 Sep 6. Dev Biol. 2010. PMID: 20804746 Free PMC article.
-
Muscleblind-like 1 is required for normal heart valve development in vivo.BMC Dev Biol. 2015 Oct 15;15:36. doi: 10.1186/s12861-015-0087-4. BMC Dev Biol. 2015. PMID: 26472242 Free PMC article.
-
Nfatc1 directs the endocardial progenitor cells to make heart valve primordium.Trends Cardiovasc Med. 2013 Nov;23(8):294-300. doi: 10.1016/j.tcm.2013.04.003. Epub 2013 May 10. Trends Cardiovasc Med. 2013. PMID: 23669445 Free PMC article. Review.
Cited by
-
Spatial transcriptomics reveals novel genes during the remodelling of the embryonic human arterial valves.PLoS Genet. 2023 Nov 27;19(11):e1010777. doi: 10.1371/journal.pgen.1010777. eCollection 2023 Nov. PLoS Genet. 2023. PMID: 38011284 Free PMC article.
-
A cellular atlas of Pitx2-dependent cardiac development.Development. 2019 Jun 14;146(12):dev180398. doi: 10.1242/dev.180398. Development. 2019. PMID: 31201182 Free PMC article.
-
BMP2 expression in the endocardial lineage is required for AV endocardial cushion maturation and remodeling.Dev Biol. 2017 Oct 1;430(1):113-128. doi: 10.1016/j.ydbio.2017.08.008. Epub 2017 Aug 6. Dev Biol. 2017. PMID: 28790014 Free PMC article.
-
Elucidating the developmental dynamics of mouse stromal cells at single-cell level.Life Med. 2022 Sep 5;1(1):45-48. doi: 10.1093/lifemedi/lnac037. eCollection 2022 Aug. Life Med. 2022. PMID: 39872159 Free PMC article. No abstract available.
-
Heart valve development: regulatory networks in development and disease.Circ Res. 2009 Aug 28;105(5):408-21. doi: 10.1161/CIRCRESAHA.109.201566. Circ Res. 2009. PMID: 19713546 Free PMC article. Review.
References
-
- Aikawa E, Nahrendorf M, Sosnovik D, Lok VM, Jaffer FA, Aikawa M, Weissleder R. Multimodality molecular imaging identifies proteolytic and osteogenic activities in early aortic valve disease. Circulation 115: 377–386, 2007. - PubMed
-
- Ameye L, Young MF. Mice deficient in small leucine-rich proteoglycans: novel in vivo models for osteoporosis, osteoarthritis, Ehlers-Danlos syndrome, muscular dystrophy, and corneal diseases. Glycobiology 12: 107R–116R, 2002. - PubMed
-
- Bergwerff M, Gittenberger-de Groot AC, Wisse LJ, DeRuiter MC, Wessels A, Martin JF, Olson EN, Kern MJ. Loss of function of the Prx1 and Prx2 homeobox genes alters architecture of the great elastic arteries and ductus arteriosus. Virchows Arch 436: 12–19, 2000. - PubMed
-
- Bialek P, Kern B, Yang X, Schrock M, Sosic D, Hong N, Wu H, Yu K, Ornitz DM, Olson EN, Justice MJ, Karsenty G. A twist code determines the onset of osteoblast differentiation. Dev Cell 6: 423–435, 2004. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

