Characterizing complex dynamics in the transactivation response element apical loop and motional correlations with the bulge by NMR, molecular dynamics, and mutagenesis
- PMID: 18621815
- PMCID: PMC2553144
- DOI: 10.1529/biophysj.108.140285
Characterizing complex dynamics in the transactivation response element apical loop and motional correlations with the bulge by NMR, molecular dynamics, and mutagenesis
Abstract
The HIV-1 transactivation response element (TAR) RNA binds a variety of proteins and is a target for developing anti-HIV therapies. TAR has two primary binding sites: a UCU bulge and a CUGGGA apical loop. We used NMR residual dipolar couplings, carbon spin relaxation (R(1) and R(2)), and relaxation dispersion (R(1rho)) in conjunction with molecular dynamics and mutagenesis to characterize the dynamics of the TAR apical loop and investigate previously proposed long-range interactions with the distant bulge. Replacement of the wild-type apical loop with a UUCG loop did not significantly affect the structural dynamics at the bulge, indicating that the apical loop and the bulge act largely as independent dynamical recognition centers. The apical loop undergoes complex dynamics at multiple timescales that are likely important for adaptive recognition: U31 and G33 undergo limited motions, G32 is highly flexible at picosecond-nanosecond timescales, and G34 and C30 form a dynamic Watson-Crick basepair in which G34 and A35 undergo a slow (approximately 30 mus) likely concerted looping in and out motion, with A35 also undergoing large amplitude motions at picosecond-nanosecond timescales. Our study highlights the power of combining NMR, molecular dynamics, and mutagenesis in characterizing RNA dynamics.
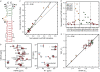
Figures



Similar articles
-
Shortening the HIV-1 TAR RNA Bulge by a Single Nucleotide Preserves Motional Modes over a Broad Range of Time Scales.Biochemistry. 2016 Aug 16;55(32):4445-56. doi: 10.1021/acs.biochem.6b00285. Epub 2016 Aug 4. Biochemistry. 2016. PMID: 27232530 Free PMC article.
-
Domain-elongation NMR spectroscopy yields new insights into RNA dynamics and adaptive recognition.RNA. 2009 Nov;15(11):1941-8. doi: 10.1261/rna.1806909. Epub 2009 Sep 23. RNA. 2009. PMID: 19776156 Free PMC article. Review.
-
Probing Na(+)-induced changes in the HIV-1 TAR conformational dynamics using NMR residual dipolar couplings: new insights into the role of counterions and electrostatic interactions in adaptive recognition.Biochemistry. 2007 Jun 5;46(22):6525-35. doi: 10.1021/bi700335n. Epub 2007 May 9. Biochemistry. 2007. PMID: 17488097 Free PMC article.
-
Base flexibility in HIV-2 TAR RNA mapped by solution (15)N, (13)C NMR relaxation.J Mol Biol. 2002 Mar 22;317(2):263-78. doi: 10.1006/jmbi.2001.5424. J Mol Biol. 2002. PMID: 11902842
-
RNA in motion.Curr Opin Chem Biol. 2008 Dec;12(6):612-8. doi: 10.1016/j.cbpa.2008.09.033. Epub 2008 Oct 26. Curr Opin Chem Biol. 2008. PMID: 18957331 Free PMC article. Review.
Cited by
-
Dynamic motions of the HIV-1 frameshift site RNA.Biophys J. 2015 Feb 3;108(3):644-54. doi: 10.1016/j.bpj.2014.12.006. Biophys J. 2015. PMID: 25650931 Free PMC article.
-
Structure of a low-population binding intermediate in protein-RNA recognition.Proc Natl Acad Sci U S A. 2016 Jun 28;113(26):7171-6. doi: 10.1073/pnas.1521349113. Epub 2016 Jun 10. Proc Natl Acad Sci U S A. 2016. PMID: 27286828 Free PMC article.
-
Rapid and accurate determination of atomistic RNA dynamic ensemble models using NMR and structure prediction.Nat Commun. 2020 Nov 2;11(1):5531. doi: 10.1038/s41467-020-19371-y. Nat Commun. 2020. PMID: 33139729 Free PMC article.
-
Molecular insights into the interaction between a disordered protein and a folded RNA.Proc Natl Acad Sci U S A. 2024 Dec 3;121(49):e2409139121. doi: 10.1073/pnas.2409139121. Epub 2024 Nov 26. Proc Natl Acad Sci U S A. 2024. PMID: 39589885 Free PMC article.
-
Shortening the HIV-1 TAR RNA Bulge by a Single Nucleotide Preserves Motional Modes over a Broad Range of Time Scales.Biochemistry. 2016 Aug 16;55(32):4445-56. doi: 10.1021/acs.biochem.6b00285. Epub 2016 Aug 4. Biochemistry. 2016. PMID: 27232530 Free PMC article.
References
-
- Muesing, M. A., D. H. Smith, and D. J. Capon. 1987. Regulation of mRNA accumulation by a human immunodeficiency virus trans-activator protein. Cell. 48:691–701. - PubMed
-
- Bannwarth, S., and A. Gatignol. 2005. HIV-1 TAR RNA: the target of molecular interactions between the virus and its host. Curr. HIV Res. 3:61–71. - PubMed
-
- Karn, J. 1999. Tackling Tat. J. Mol. Biol. 293:235–254. - PubMed
-
- Jones, K. A. 1997. Taking a new TAK on Tat transactivation. Genes Dev. 11:2593–2599. - PubMed
-
- Majello, B., G. Napolitano, A. Giordano, and L. Lania. 1999. Transcriptional regulation by targeted recruitment of cyclin-dependent CDK9 kinase in vivo. Oncogene. 18:4598–4605. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

