Mutational analysis of the connector segment in the HAMP domain of Tsr, the Escherichia coli serine chemoreceptor
- PMID: 18621896
- PMCID: PMC2566183
- DOI: 10.1128/JB.00750-08
Mutational analysis of the connector segment in the HAMP domain of Tsr, the Escherichia coli serine chemoreceptor
Abstract
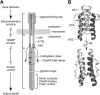
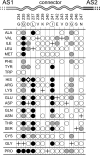
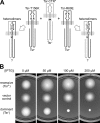
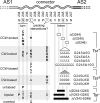
HAMP domains are approximately 50-residue motifs, found in many bacterial signaling proteins, that consist of two amphiphilic helices joined by a nonhelical connector segment. The HAMP domain of Tsr, the serine chemoreceptor of Escherichia coli, receives transmembrane input signals from the periplasmic serine binding domain and in turn modulates output signals from the Tsr kinase control domain to elicit chemotactic responses. We created random amino acid replacements at each of the 14 connector residues of Tsr-HAMP to identify those that are critical for Tsr function. In all, we surveyed 179 connector missense mutants and identified three critical residues (G235, L237, and I241) at which most replacements destroyed Tsr function and another important residue (G245) at which most replacements impaired Tsr function. The region surrounding G245 tolerated 1-residue deletions and insertions of up to 10 glycines, suggesting a role as a relatively nonspecific, flexible linker. The critical connector residues are consistent with a structural model of the Tsr-HAMP domain based on the solution structure of an isolated thermophile HAMP domain (M. Hulko, F. Berndt, M. Gruber, J. U. Linder, V. Truffault, A. Schultz, J. Martin, J. E. Schultz, A. N. Lupas, and M. Coles, Cell 126:929-940, 2006) in which G235 defines a critical turn at the C terminus of the first helix and L237 and I241 pack against the helices, perhaps to stabilize alternative HAMP signaling conformations. Most I241 lesions locked Tsr signal output in the kinase-on mode, implying that this residue is responsible mainly for stabilizing the kinase-off signaling state. In contrast, lesions at L237 resulted in a variety of aberrant output patterns, suggesting a role in toggling output between signaling states.
Figures


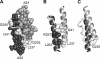
References
-
- Ames, P., and J. S. Parkinson. 1988. Transmembrane signaling by bacterial chemoreceptors: E. coli transducers with locked signal output. Cell 55817-826. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

