Structural comparisons of class I phosphoinositide 3-kinases
- PMID: 18633356
- PMCID: PMC2847604
- DOI: 10.1038/nrc2443
Structural comparisons of class I phosphoinositide 3-kinases
Abstract
Class I phosphoinositide 3-kinases (PI3Ks) are lipid kinases that regulate cell growth. One of these kinases, PI3Kalpha, is frequently mutated in diverse tumour types. The recently determined structure of PI3Kalpha reveals features that distinguish this enzyme from related lipid kinases. In addition, wild-type PI3Kgamma differs from PI3Kalpha by a substitution identical to a PI3Kalpha oncogenic mutant (His1047Arg) that might explain the differences in the enzymatic activities of the normal and mutant PI3Kalpha. Comparison of the PI3K structures also identified structural features that could potentially be exploited for the design of isoform-specific inhibitors.
Figures


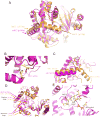
References
-
- Manning BD, Tee AR, Logsdon MN, Blenis J, Cantley LC. Identification of the tuberous sclerosis complex-2 tumor suppressor gene product tuberin as a target of the phosphoinositide 3-kinase/akt pathway. Mol Cell. 2002;10:151–62. - PubMed
-
- Cross DA, Alessi DR, Cohen P, Andjelkovich M, Hemmings BA. Inhibition of glycogen synthase kinase-3 by insulin mediated by protein kinase B. Nature. 1995;378:785–9. - PubMed
-
- Datta SR, et al. Akt phosphorylation of BAD couples survival signals to the cell-intrinsic death machinery. Cell. 1997;91:231–41. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

