Role of tRNA-like structures in controlling plant virus replication
- PMID: 18638511
- PMCID: PMC2676847
- DOI: 10.1016/j.virusres.2008.06.010
Role of tRNA-like structures in controlling plant virus replication
Abstract
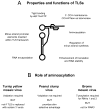
Transfer RNA-like structures (TLSs) that are sophisticated functional mimics of tRNAs are found at the 3'-termini of the genomes of a number of plant positive strand RNA viruses. Three natural aminoacylation identities are represented: valine, histidine, and tyrosine. Paralleling this variety in structure, the roles of TLSs vary widely between different viruses. For Turnip yellow mosaic virus, the TLS must be capable of valylation in order to support infectivity, major roles being the provision of translational enhancement and down-regulation of minus strand initiation. In contrast, valylation of the Peanut clump virus TLS is not essential. An intermediate situation seems to exist for Brome mosaic virus, whose RNAs 1 and 2, but not RNA 3, need to be capable of tyrosylation to support infectivity. Other known roles for certain TLSs include: (i) the recruitment of host CCA nucleotidyltransferase as a telomerase to maintain intact 3' CCA termini, (ii) involvement in the encapsidation of viral RNAs, and (iii) presentation of minus strand promoter elements for replicase recognition. In the latter role, the promoter elements reside within the TLS but are not functionally dependent on tRNA mimicry. The phylogenetic distribution of TLSs indicates that their evolutionary history includes frequent horizontal exchange, as has been observed for protein-coding regions of plant positive strand RNA viruses.
Figures




Similar articles
-
Transfer RNA mimicry in a new group of positive-strand RNA plant viruses, the furoviruses: differential aminoacylation between the RNA components of one genome.Virology. 1998 Jun 20;246(1):170-8. doi: 10.1006/viro.1998.9193. Virology. 1998. PMID: 9657004
-
Comparison and functional implications of the 3D architectures of viral tRNA-like structures.RNA. 2009 Feb;15(2):294-307. doi: 10.1261/rna.1360709. RNA. 2009. PMID: 19144910 Free PMC article.
-
The turnip yellow mosaic virus tRNA-like structure cannot be replaced by generic tRNA-like elements or by heterologous 3' untranslated regions known to enhance mRNA expression and stability.J Virol. 1996 Apr;70(4):2107-15. doi: 10.1128/JVI.70.4.2107-2115.1996. J Virol. 1996. PMID: 8642631 Free PMC article.
-
Viral tRNAs and tRNA-like structures.Wiley Interdiscip Rev RNA. 2010 Nov-Dec;1(3):402-14. doi: 10.1002/wrna.42. Epub 2010 Sep 13. Wiley Interdiscip Rev RNA. 2010. PMID: 21956939 Review.
-
Cis-acting RNA elements in positive-strand RNA plant virus genomes.Virology. 2015 May;479-480:434-43. doi: 10.1016/j.virol.2015.02.032. Epub 2015 Mar 7. Virology. 2015. PMID: 25759098 Review.
Cited by
-
The tRNA-like small noncoding RNA mascRNA promotes global protein translation.EMBO Rep. 2020 Dec 3;21(12):e49684. doi: 10.15252/embr.201949684. Epub 2020 Oct 19. EMBO Rep. 2020. PMID: 33073493 Free PMC article.
-
Molecular mimicry of human tRNALys anti-codon domain by HIV-1 RNA genome facilitates tRNA primer annealing.RNA. 2013 Feb;19(2):219-29. doi: 10.1261/rna.036681.112. Epub 2012 Dec 21. RNA. 2013. PMID: 23264568 Free PMC article.
-
Differential Expression of Genes between a Tolerant and a Susceptible Maize Line in Response to a Sugarcane Mosaic Virus Infection.Viruses. 2022 Aug 17;14(8):1803. doi: 10.3390/v14081803. Viruses. 2022. PMID: 36016425 Free PMC article.
-
Identification of potential conserved RNA secondary structure throughout influenza A coding regions.RNA. 2011 Jun;17(6):991-1011. doi: 10.1261/rna.2619511. Epub 2011 May 2. RNA. 2011. PMID: 21536710 Free PMC article.
-
Identification and Characterization of a Class of MALAT1-like Genomic Loci.Cell Rep. 2017 May 23;19(8):1723-1738. doi: 10.1016/j.celrep.2017.05.006. Cell Rep. 2017. PMID: 28538188 Free PMC article.
References
-
- Agranovsky AA, Dolja VV, Gorbulev VG, Kozlov YV, Atabekov JG. Aminoacylation of barley stripe mosaic virus RNA: polyadenylate-containing RNA has a 3′-terminal tyrosine-accepting structure. Virology. 1981;113:174–187. - PubMed
-
- Agranovsky AA, Karasev AV, Novikov VK, Lunina NA, Loginov S, Tyulkina LG. Poa semilatent virus, a hordeivirus having no internal polydisperse poly(A) in the 3′ non-coding region of the RNA genome. J Gen Virol. 1992;73:2085–92. - PubMed
-
- Ahlquist P, Dasgupta R, Kaesberg P. Near identity of 3′-RNA secondary structure in bromoviruses and cucumber mosaic virus. Cell. 1981;23:183–9. - PubMed
-
- Annamalai P, Rao AL. Dispensability of 3′ tRNA-like sequence for packaging cowpea chlorotic mottle virus genomic RNAs. Virology. 2005;332:650–8. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

