Platypus globin genes and flanking loci suggest a new insertional model for beta-globin evolution in birds and mammals
- PMID: 18657265
- PMCID: PMC2529266
- DOI: 10.1186/1741-7007-6-34
Platypus globin genes and flanking loci suggest a new insertional model for beta-globin evolution in birds and mammals
Abstract
Background: Vertebrate alpha (alpha)- and beta (beta)-globin gene families exemplify the way in which genomes evolve to produce functional complexity. From tandem duplication of a single globin locus, the alpha- and beta-globin clusters expanded, and then were separated onto different chromosomes. The previous finding of a fossil beta-globin gene (omega) in the marsupial alpha-cluster, however, suggested that duplication of the alpha-beta cluster onto two chromosomes, followed by lineage-specific gene loss and duplication, produced paralogous alpha- and beta-globin clusters in birds and mammals. Here we analyse genomic data from an egg-laying monotreme mammal, the platypus (Ornithorhynchus anatinus), to explore haemoglobin evolution at the stem of the mammalian radiation.
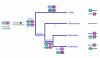
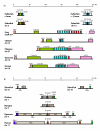
Results: The platypus alpha-globin cluster (chromosome 21) contains embryonic and adult alpha- globin genes, a beta-like omega-globin gene, and the GBY globin gene with homology to cytoglobin, arranged as 5'-zeta-zeta'-alphaD-alpha3-alpha2-alpha1-omega-GBY-3'. The platypus beta-globin cluster (chromosome 2) contains single embryonic and adult globin genes arranged as 5'-epsilon-beta-3'. Surprisingly, all of these globin genes were expressed in some adult tissues. Comparison of flanking sequences revealed that all jawed vertebrate alpha-globin clusters are flanked by MPG-C16orf35 and LUC7L, whereas all bird and mammal beta-globin clusters are embedded in olfactory genes. Thus, the mammalian alpha- and beta-globin clusters are orthologous to the bird alpha- and beta-globin clusters respectively.
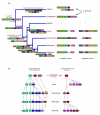
Conclusion: We propose that alpha- and beta-globin clusters evolved from an ancient MPG-C16orf35-alpha-beta-GBY-LUC7L arrangement 410 million years ago. A copy of the original beta (represented by omega in marsupials and monotremes) was inserted into an array of olfactory genes before the amniote radiation (>315 million years ago), then duplicated and diverged to form orthologous clusters of beta-globin genes with different expression profiles in different lineages.
Figures


Similar articles
-
Globin genes on the move.J Biol. 2008 Nov 20;7(9):35. doi: 10.1186/jbiol92. J Biol. 2008. PMID: 19040770 Free PMC article. Review.
-
Globin gene structure in a reptile supports the transpositional model for amniote α- and β-globin gene evolution.Chromosome Res. 2010 Dec;18(8):897-907. doi: 10.1007/s10577-010-9164-5. Epub 2010 Nov 30. Chromosome Res. 2010. PMID: 21116705
-
Sequencing and mapping hemoglobin gene clusters in the Australian model dasyurid marsupial Sminthopsis macroura.Cytogenet Genome Res. 2005;108(4):333-41. doi: 10.1159/000081528. Cytogenet Genome Res. 2005. PMID: 15627754
-
Linkage of the beta-like omega-globin gene to alpha-like globin genes in an Australian marsupial supports the chromosome duplication model for separation of globin gene clusters.J Mol Evol. 2004 Jun;58(6):642-52. doi: 10.1007/s00239-004-2584-0. J Mol Evol. 2004. PMID: 15461421
-
Use of long sequence alignments to study the evolution and regulation of mammalian globin gene clusters.Mol Biol Evol. 1993 Jan;10(1):73-102. doi: 10.1093/oxfordjournals.molbev.a039991. Mol Biol Evol. 1993. PMID: 8383794 Review.
Cited by
-
Globin genes on the move.J Biol. 2008 Nov 20;7(9):35. doi: 10.1186/jbiol92. J Biol. 2008. PMID: 19040770 Free PMC article. Review.
-
Evolution of hemoglobin and its genes.Cold Spring Harb Perspect Med. 2012 Dec 1;2(12):a011627. doi: 10.1101/cshperspect.a011627. Cold Spring Harb Perspect Med. 2012. PMID: 23209182 Free PMC article.
-
A conserved cluster of three PRD-class homeobox genes (homeobrain, rx and orthopedia) in the Cnidaria and Protostomia.Evodevo. 2010 Jul 5;1(1):3. doi: 10.1186/2041-9139-1-3. Evodevo. 2010. PMID: 20849646 Free PMC article.
-
The globin gene family of the cephalochordate amphioxus: implications for chordate globin evolution.BMC Evol Biol. 2010 Nov 30;10:370. doi: 10.1186/1471-2148-10-370. BMC Evol Biol. 2010. PMID: 21118516 Free PMC article.
-
Developmental regulation of hemoglobin synthesis in the green anole lizard Anolis carolinensis.J Exp Biol. 2011 Feb 15;214(Pt 4):575-81. doi: 10.1242/jeb.050443. J Exp Biol. 2011. PMID: 21270305 Free PMC article.
References
-
- Dickerson RE, Geis I. Hemoglobin: Structure, Function, Evolution and Pathology. Menlo Park, CA: Benjamin Cummings; 1983. pp. 65–116.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

