HLA-C increases HIV-1 infectivity and is associated with gp120
- PMID: 18673537
- PMCID: PMC2531131
- DOI: 10.1186/1742-4690-5-68
HLA-C increases HIV-1 infectivity and is associated with gp120
Abstract
Background: A recently identified genetic polymorphism located in the 5' region of the HLA-C gene is associated with individual variations in HIV-1 viral load and with differences in HLA-C expression levels. HLA-C has the potential to restrict HIV-1 by presenting epitopes to cytotoxic T cells but it is also a potent inhibitor of NK cells. In addition, HLA-C molecules incorporated within the HIV-1 envelope have been shown to bind to the envelope glycoprotein gp120 and enhance viral infectivity. We investigated this last property in cell fusion assays where the expression of HLA-C was silenced by small interfering RNA sequences. Syncytia formation was analyzed by co-cultivating cell lines expressing HIV-1 gp120/gp41 from different laboratory and primary isolates with target cells expressing different HIV-1 co-receptors. Virus infectivity was analyzed using pseudoviruses. Molecular complexes generated during cell fusion (fusion complexes) were purified and analyzed for their HLA-C content.
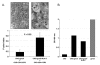
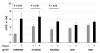
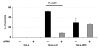
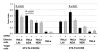
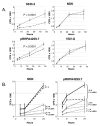
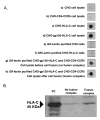
Results: HLA-C positive cells co-expressing HIV-1 gp120/gp41 fused more rapidly and produced larger syncytia than HLA-C negative cells. Transient transfection of gp120/gp41 from different primary isolates in HLA-C positive cells resulted in a significant cell fusion increase. Fusion efficiency was reduced in HLA-C silenced cells compared to non-silenced cells when co-cultivated with different target cell lines expressing HIV-1 co-receptors. Similarly, pseudoviruses produced from HLA-C silenced cells were significantly less infectious. HLA-C was co-purified with gp120 from cells before and after fusion and was associated with the fusion complex.
Conclusion: Virionic HLA-C molecules associate to Env and increase the infectivity of both R5 and X4 viruses. Genetic polymorphisms associated to variations in HLA-C expression levels may therefore influence the individual viral set point not only by means of a regulation of the virus-specific immune response but also via a direct effect on the virus replicative capacity. These findings have implications for the understanding of the HIV-1 entry mechanism and of the role of Env conformational modifications induced by virion-associated host proteins.
Figures

Similar articles
-
The Polar Region of the HIV-1 Envelope Protein Determines Viral Fusion and Infectivity by Stabilizing the gp120-gp41 Association.J Virol. 2019 Mar 21;93(7):e02128-18. doi: 10.1128/JVI.02128-18. Print 2019 Apr 1. J Virol. 2019. PMID: 30651369 Free PMC article.
-
Enhanced HIV infectivity and changes in GP120 conformation associated with viral incorporation of human leucocyte antigen class I molecules.AIDS. 1999 Oct 22;13(15):2033-42. doi: 10.1097/00002030-199910220-00005. AIDS. 1999. PMID: 10546855
-
Mutations That Increase the Stability of the Postfusion gp41 Conformation of the HIV-1 Envelope Glycoprotein Are Selected by both an X4 and R5 HIV-1 Virus To Escape Fusion Inhibitors Corresponding to Heptad Repeat 1 of gp41, but the gp120 Adaptive Mutations Differ between the Two Viruses.J Virol. 2019 May 15;93(11):e00142-19. doi: 10.1128/JVI.00142-19. Print 2019 Jun 1. J Virol. 2019. PMID: 30894471 Free PMC article.
-
CD4 activation of HIV fusion.Int J Cell Cloning. 1992 Nov;10(6):323-32. doi: 10.1002/stem.5530100603. Int J Cell Cloning. 1992. PMID: 1281202 Review.
-
HIV-envelope-dependent cell-cell fusion: quantitative studies.ScientificWorldJournal. 2009 Aug 11;9:746-63. doi: 10.1100/tsw.2009.90. ScientificWorldJournal. 2009. PMID: 19705036 Free PMC article. Review.
Cited by
-
Stability and Expression Levels of HLA-C on the Cell Membrane Modulate HIV-1 Infectivity.J Virol. 2017 Dec 14;92(1):e01711-17. doi: 10.1128/JVI.01711-17. Print 2018 Jan 1. J Virol. 2017. PMID: 29070683 Free PMC article.
-
HIV-1-Associated Neurocognitive Disorders: Is HLA-C Binding Stability to β2-Microglobulin a Missing Piece of the Pathogenetic Puzzle?Front Neurol. 2018 Sep 21;9:791. doi: 10.3389/fneur.2018.00791. eCollection 2018. Front Neurol. 2018. PMID: 30298049 Free PMC article.
-
HIV-1 Env associates with HLA-C free-chains at the cell membrane modulating viral infectivity.Sci Rep. 2017 Jan 4;7:40037. doi: 10.1038/srep40037. Sci Rep. 2017. PMID: 28051183 Free PMC article.
-
Development of a platelet-activating factor antagonist for HIV-1 associated neurocognitive disorders.J Neuroimmunol. 2009 Aug 18;213(1-2):47-59. doi: 10.1016/j.jneuroim.2009.06.002. Epub 2009 Jun 21. J Neuroimmunol. 2009. PMID: 19541372 Free PMC article.
-
Convergent Evolution of HLA-C Downmodulation in HIV-1 and HIV-2.mBio. 2020 Jul 14;11(4):e00782-20. doi: 10.1128/mBio.00782-20. mBio. 2020. PMID: 32665270 Free PMC article.
References
-
- Fellay J, Shianna KV, Ge D, Colombo S, Ledergerber B, Weale M, Zhang K, Gumbs C, Castagna A, Cossarizza A, Cozzi-Lepri A, De Luca A, Easterbrook P, Francioli P, Mallal S, Martinez-Picado J, Miro JM, Obel N, Smith JP, Wyniger J, Descombes P, Antonarakis SE, Letvin NL, McMichael AJ, Haynes BF, Telenti A, Goldstein DB. Science. 2007/07/21. Vol. 317. 2007. A whole-genome association study of major determinants for host control of HIV-1; pp. 944–947. - DOI - PMC - PubMed
-
- Goulder PJ, Edwards A, Phillips RE, McMichael AJ. Aids. 1997/12/31. Vol. 11. 1997. Identification of a novel HLA-A24-restricted cytotoxic T-lymphocyte epitope within HIV-1 Nef; pp. 1883–1884. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

