Gene expression analysis of host innate immune responses during Lethal H5N1 infection in ferrets
- PMID: 18684821
- PMCID: PMC2573250
- DOI: 10.1128/JVI.00691-08
Gene expression analysis of host innate immune responses during Lethal H5N1 infection in ferrets
Abstract
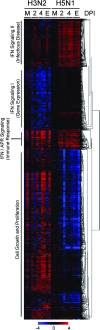
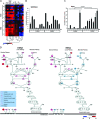
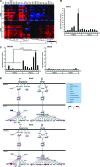
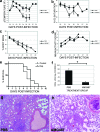
How viral and host factors contribute to the severe pathogenicity of the H5N1 subtype of avian influenza virus infection in humans is poorly understood. We identified three clusters of differentially expressed innate immune response genes in lungs from H5N1 (A/Vietnam/1203/04) influenza virus-infected ferrets by oligonucleotide microarray analysis. Interferon response genes were more strongly expressed in H5N1-infected ferret lungs than in lungs from ferrets infected with the less pathogenic H3N2 subtype. In particular, robust CXCL10 gene expression in H5N1-infected ferrets led us to test the pathogenic role of signaling via CXCL10's cognate receptor, CXCR3, during H5N1 influenza virus infection. Treatment of H5N1-infected ferrets with the drug AMG487, a CXCR3 antagonist, resulted in a reduction of symptom severity and delayed mortality compared to vehicle treatment. We contend that unregulated host interferon responses are at least partially responsible for the severity of H5N1 infection and provide evidence that attenuating the CXCR3 signaling pathway improves the clinical course of H5N1 infection in ferrets.
Figures
References
-
- Abele, R., and R. Tampe. 2004. The ABCs of immunology: structure and function of TAP, the transporter associated with antigen processing. Physiology (Bethesda) 19216-224. - PubMed
-
- Beebe, D. P., R. D. Schreiber, and N. R. Cooper. 1983. Neutralization of influenza virus by normal human sera: mechanisms involving antibody and complement. J. Immunol. 1301317-1322. - PubMed
-
- Bjornson, A. B., M. A. Mellencamp, and G. M. Schiff. 1991. Complement is activated in the upper respiratory tract during influenza virus infection. Am. Rev. Respir. Dis. 1431062-1066. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

