Acquisition of classical CTX prophage from Vibrio cholerae O141 by El Tor strains aided by lytic phages and chitin-induced competence
- PMID: 18689675
- PMCID: PMC2575248
- DOI: 10.1073/pnas.0805560105
Acquisition of classical CTX prophage from Vibrio cholerae O141 by El Tor strains aided by lytic phages and chitin-induced competence
Abstract
The El Tor biotype of Vibrio cholerae O1, causing the current seventh pandemic of cholera, has replaced the classical biotype, which caused the sixth pandemic. The CTX prophages encoding cholera toxin in the two biotypes have distinct repressor (rstR) genes. Recently, new variants of El Tor strains that carry the classical type (CTX(class)) prophage have emerged. These "hybrid" strains apparently originate through lateral gene transfer and recombination events. To explore possible donors of the CTX(class) prophage and its mode of transfer, we tested environmental V. cholerae isolates for the presence of CTX(class) prophage and mobility of the phage genome. Of the 272 environmental V. cholerae isolates tested, 6 were found to carry the CTX(class) prophage; all of these belonged to the O141 serogroup. These O141 strains were unable to produce infectious CTX(class) phage or to transmit the prophage to recipient strains in the mouse model of infection; however, the CTX(class) prophage was acquired by El Tor strains when cultured with the O141 strains in microcosms composed of filtered environmental water, a chitin substrate, and a V. cholerae O141-specific bacteriophage. The CTX(class) prophage either coexisted with or replaced the resident CTX(ET) prophage, resulting in El Tor strains with CTX genotypes similar to those of the naturally occurring hybrid strains. Our results support a model involving phages and natural chitin substrate in the emergence of new variants of pathogenic V. cholerae. Furthermore, the O141 strains apparently represent an alternative reservoir of the CTX(class) phage genome, because the classical V. cholerae O1 strains are possibly extinct.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

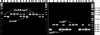

Similar articles
-
Genomic analysis of the Mozambique strain of Vibrio cholerae O1 reveals the origin of El Tor strains carrying classical CTX prophage.Proc Natl Acad Sci U S A. 2007 Mar 20;104(12):5151-6. doi: 10.1073/pnas.0700365104. Epub 2007 Mar 12. Proc Natl Acad Sci U S A. 2007. PMID: 17360342 Free PMC article.
-
Molecular insights into the evolutionary pathway of Vibrio cholerae O1 atypical El Tor variants.PLoS Pathog. 2014 Sep 18;10(9):e1004384. doi: 10.1371/journal.ppat.1004384. eCollection 2014 Sep. PLoS Pathog. 2014. PMID: 25233006 Free PMC article.
-
Genesis of variants of Vibrio cholerae O1 biotype El Tor: role of the CTXphi array and its position in the genome.Microbiology (Reading). 2003 Jan;149(Pt 1):89-97. doi: 10.1099/mic.0.25599-0. Microbiology (Reading). 2003. PMID: 12576583
-
CTX prophages in Vibrio cholerae O1 strains.J Microbiol Biotechnol. 2014 Jun 28;24(6):725-31. doi: 10.4014/jmb.1403.03063. J Microbiol Biotechnol. 2014. PMID: 24722374 Review.
-
Revisiting the Global Epidemiology of Cholera in Conjuction With the Genomics of Vibrio cholerae.Front Public Health. 2019 Jul 23;7:203. doi: 10.3389/fpubh.2019.00203. eCollection 2019. Front Public Health. 2019. PMID: 31396501 Free PMC article. Review.
Cited by
-
Non-coding sRNAs regulate virulence in the bacterial pathogen Vibrio cholerae.RNA Biol. 2012 Apr;9(4):392-401. doi: 10.4161/rna.19975. Epub 2012 Apr 1. RNA Biol. 2012. PMID: 22546941 Free PMC article. Review.
-
CTXφ Replication Depends on the Histone-Like HU Protein and the UvrD Helicase.PLoS Genet. 2015 May 20;11(5):e1005256. doi: 10.1371/journal.pgen.1005256. eCollection 2015 May. PLoS Genet. 2015. PMID: 25992634 Free PMC article.
-
Molecular keys of the tropism of integration of the cholera toxin phage.Proc Natl Acad Sci U S A. 2010 Mar 2;107(9):4377-82. doi: 10.1073/pnas.0910212107. Epub 2010 Jan 21. Proc Natl Acad Sci U S A. 2010. PMID: 20133778 Free PMC article.
-
Pandemic Vibrio cholerae shuts down site-specific recombination to retain an interbacterial defence mechanism.Nat Commun. 2020 Dec 7;11(1):6246. doi: 10.1038/s41467-020-20012-7. Nat Commun. 2020. PMID: 33288753 Free PMC article.
-
Conversion of a recA-Mediated Non-toxigenic Vibrio cholerae O1 Strain to a Toxigenic Strain Using Chitin-Induced Transformation.Front Microbiol. 2019 Nov 7;10:2562. doi: 10.3389/fmicb.2019.02562. eCollection 2019. Front Microbiol. 2019. PMID: 31787954 Free PMC article.
References
-
- Waldor MK, Mekalanos JJ. Lysogenic conversion by a filamentous bacteriophage encoding cholera toxin. Science. 1996;272:1910–1914. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

