Relationship between homoeologous regulatory and structural genes in allopolyploid genome - a case study in bread wheat
- PMID: 18700978
- PMCID: PMC2538534
- DOI: 10.1186/1471-2229-8-88
Relationship between homoeologous regulatory and structural genes in allopolyploid genome - a case study in bread wheat
Abstract
Background: The patterns of expression of homoeologous genes in hexaploid bread wheat have been intensively studied in recent years, but the interaction between structural genes and their homoeologous regulatory genes remained unclear. The question was as to whether, in an allopolyploid, this interaction is genome-specific, or whether regulation cuts across genomes. The aim of the present study was cloning, sequence analysis, mapping and expression analysis of F3H (flavanone 3-hydroxylase - one of the key enzymes in the plant flavonoid biosynthesis pathway) homoeologues in bread wheat and study of the interaction between F3H and their regulatory genes homoeologues - Rc (red coleoptiles).
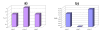
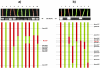
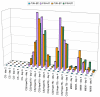
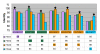
Results: PCR-based cloning of F3H sequences from hexaploid bread wheat (Triticum aestivum L.), a wild tetraploid wheat (T. timopheevii) and their putative diploid progenitors was employed to localize, physically map and analyse the expression of four distinct bread wheat F3H copies. Three of these form a homoeologous set, mapping to the chromosomes of homoeologous group 2; they are highly similar to one another at the structural and functional levels. However, the fourth copy is less homologous, and was not expressed in anthocyanin pigmented coleoptiles. The presence of dominant alleles at the Rc-1 homoeologous loci, which are responsible for anthocyanin pigmentation in the coleoptile, was correlated with F3H expression in pigmented coleoptiles. Each dominant Rc-1 allele affected the expression of the three F3H homoeologues equally, but the level of F3H expression was dependent on the identity of the dominant Rc-1 allele present. Thus, the homoeologous Rc-1 genes contribute more to functional divergence than do the structural F3H genes.
Conclusion: The lack of any genome-specific relationship between F3H-1 and Rc-1 implies an integrative evolutionary process among the three diploid genomes, following the formation of hexaploid wheat. Regulatory genes probably contribute more to the functional divergence between the wheat genomes than do the structural genes themselves. This is in line with the growing consensus which suggests that although heritable morphological traits are determined by the expression of structural genes, it is the regulatory genes which are the prime determinants of allelic identity.
Figures






References
-
- Bogdanova ED, Sarbaev AT, Makhmudova KK. Resistance of common wheat to bunt [abstract] Proceedings of the Research Conference on Genetics: 2002; Moscow. 2002. pp. 43–44.
-
- Ryan KG, Swinny EE, Winefield C, Markham KR. Flavonoids and UV photoprotection in Arabidopsis mutants. Z Naturforsch [C] 2001;56:745–754. - PubMed
-
- Khlestkina EK, Röder MS, Pshenichnikova TA, Simonov AV, Salina EA, Börner A. Genes for anthocyanin pigmentation in wheat: review and microsatellite-based mapping. In: Verrity JF, Abbington LE, editor. Chromosome Mapping Research Developments. New York: NOVA Science Publishers, Inc, USA; 2008. pp. 155–175.https://www.novapublishers.com/catalog/product_info.php?products_id=6838
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

