Crystal structures of C4-dicarboxylate ligand complexes with sensor domains of histidine kinases DcuS and DctB
- PMID: 18701447
- PMCID: PMC2573060
- DOI: 10.1074/jbc.M805253200
Crystal structures of C4-dicarboxylate ligand complexes with sensor domains of histidine kinases DcuS and DctB
Abstract
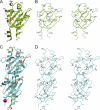
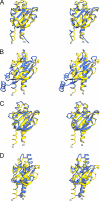
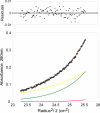
Two-component signaling systems allow bacteria to adapt to changing environments. Typically, a chemical or other stimulus is detected by the periplasmic sensor domain of a transmembrane histidine kinase sensor, which in turn relays a signal through a phosphotransfer cascade to the cognate cytoplasmic response regulator. Such systems lead ultimately to changes in gene expression or cell motility. Mechanisms of ligand binding and signal transduction through the cell membrane in histidine kinases are not fully understood. In an effort to further understand such processes, we have solved the crystal structures of the periplasmic sensor domains of Escherichia coli DcuS and of Vibrio cholerae DctB in complex with the respective cognate ligands, malate and succinate. Both proteins are involved in the regulation of the transport and metabolism of C(4)-dicarboxylates, but they are not highly related by sequence similarity. Our work reveals that despite disparate sizes, both structures contain a similar characteristic alpha/beta PDC (PhoQ-DcuS-CitA) sensor-domain fold and display similar modes of ligand binding, suggesting similar mechanisms of function.
Figures



References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

