Conformational preferences of substrates for human prolyl 4-hydroxylase
- PMID: 18702512
- PMCID: PMC2810141
- DOI: 10.1021/bi8009373
Conformational preferences of substrates for human prolyl 4-hydroxylase
Abstract
Prolyl 4-hydroxylase (P4H) catalyzes the posttranslational hydroxylation of (2 S)-proline (Pro) residues in procollagen strands. The resulting (2 S,4 R)-4-hydroxyproline (Hyp) residues are essential for the folding, secretion, and stability of the collagen triple helix. Even though its product (Hyp) differs from its substrate (Pro) by only a single oxygen atom, no product inhibition has been observed for P4H. Here, we examine the basis for the binding and turnover of substrates by human P4H. Synthetic peptides containing (2 S,4 R)-4-fluoroproline (Flp), (2 S,4 S)-4-fluoroproline (flp), (2 S)-4-ketoproline (Kep), (2 S)-4-thiaproline (Thp), and 3,5-methanoproline (Mtp) were evaluated as substrates for P4H. Peptides containing Pro, flp, and Thp were found to be excellent substrates for P4H, forming Hyp, Kep, and (2 S,4 R)-thiaoxoproline, respectively. Thus, P4H is tolerant to some substitutions on C-4 of the pyrrolidine ring. In contrast, peptides containing Flp, Kep, or Mtp did not even bind to the active site of P4H. Each proline analogue that does bind to P4H is also a substrate, indicating that discrimination occurs at the level of binding rather than turnover. As the iron(IV)-oxo species that forms in the active site of P4H is highly reactive, P4H has an imperative for forming a snug complex with its substrate and appears to do so. Most notably, those proline analogues with a greater preference for a C (gamma)- endo pucker and cis peptide bond were the ones recognized by P4H. As Hyp has a strong preference for C (gamma)- exo pucker and trans peptide bond, P4H appears to discriminate against the conformation of proline residues in a manner that diminishes product inhibition during collagen biosynthesis.
Figures
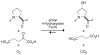







References
-
- Prockop DJ, Kivirikko KI. Collagens: Molecular biology, diseases, and potentials for therapy. Annu Rev Biochem. 1995;64:403–434. - PubMed
-
- Ramshaw JAM, Shah NK, Brodsky B. Gly–X–Y tripeptide frequencies in collagen: A context for host–guest triple-helical peptides. J Struct Biol. 1998;122:86–91. - PubMed
-
- McCaldon P, Argos P. Oligopeptide biases in protein sequences and their use in predicting protein coding regions in nucleotide sequences. Proteins: Struct Funct Genet. 1988;4:99–122. - PubMed
-
- Berg RA, Prockop DJ. The thermal transition of a non-hydroxylated form of collagen. Evidence for a role for hydroxyproline in stabilizing the triple-helix of collagen. Biochem Biophys Res Commun. 1973;52:115–120. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

