Allostery: an illustrated definition for the 'second secret of life'
- PMID: 18706817
- PMCID: PMC2574622
- DOI: 10.1016/j.tibs.2008.05.009
Allostery: an illustrated definition for the 'second secret of life'
Abstract
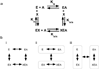
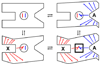
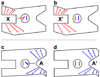
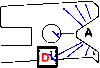
Although allosteric regulation is the 'second secret of life', the molecular mechanisms that give rise to allostery currently elude understanding. In my opinion, experimental progress is hampered by a commonly used but misleading definition of allostery as protein structural changes that are elicited by the binding of a single ligand. Allostery is more strictly defined in functional terms as a comparison of how one ligand binds in the absence, versus the presence, of a second ligand. Therefore, as each of the two binding events involves two protein complexes, a study of allostery must consider four complexes and not just two. Such a comparison can distinguish allosteric from non-allosteric protein changes, the importance of which is frequently overlooked. When a study of all four complexes is not feasible, an alternative, albeit limited, strategy can identify subsets of allosteric-specific changes.
Figures
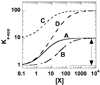
References
-
- Monod J. Chance and Necessity: Essay on the Natural Philosophy of Modern Biology. Penguin Books Ltd; 1977.
-
- Monod J, et al. Allosteric proteins and cellular control systems. J Mol Biol. 1963;6:306–329. - PubMed
-
- Groebe DR. Screening for positive allosteric modulators of biological targets. Drug discovery today. 2006;11:632–639. - PubMed
-
- May LT, et al. Allosteric modulation of G protein-coupled receptors. Annual review of pharmacology and toxicology. 2007;47:1–51. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

