Single mutations change CYP2F3 from a dehydrogenase of 3-methylindole to an oxygenase
- PMID: 18717595
- PMCID: PMC2843524
- DOI: 10.1021/bi8005658
Single mutations change CYP2F3 from a dehydrogenase of 3-methylindole to an oxygenase
Abstract
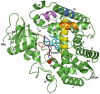
Pulmonary cytochrome P450 2F3 (CYP2F3) catalyzes the dehydrogenation of the pneumotoxin 3-methylindole (3MI) to an electrophilic intermediate, 3-methyleneindolenine, which is responsible for the toxicity of the parent compound. Members of the CYP2F subfamily are the only enzymes known to exclusively dehydrogenate 3MI, without detectable formation of oxygenation products. Thus, CYP2F3 is an attractive model to study dehydrogenation mechanisms. The purpose of this study was to identify specific residues that could facilitate 3MI dehydrogenation. Both single and double mutations were constructed to study the molecular mechanisms that direct dehydrogenation. Double mutations in substrate recognition sites (SRS) 1 produced an inactive enzyme, while double mutants in SRS 4 did not alter 3MI metabolism. However, double mutations in SRS 5 and SRS 6 successfully introduced oxygenase activity to CYP2F3. Single mutations in SRS 5, SRS 6 and near SRS 2 also introduced 3MI oxygenase activity. Mutants S474H and D361T oxygenated 3MI but also increased dehydrogenation rates, while G214L, E215Q and S475I catalyzed 3MI oxygenation exclusively. A homology model of CYP2F3 was precisely consistent with specific dehydrogenation of 3MI via initial hydrogen atom abstraction from the methyl group. In addition, intramolecular kinetic deuterium isotope studies demonstrated an isotope effect ( K H/ K D) of 6.8. This relatively high intramolecular deuterium isotope effect confirmed the initial hydrogen abstraction step; a mutant (D361T) that retained the dehydrogenation reaction exhibited the same deuterium isotope effect. The results showed that a single alteration, such as a serine to isoleucine change at residue 475, dramatically switched catalytic preference from dehydrogenation to oxygenation.
Figures




Similar articles
-
Cloning and expression of CYP2F3, a cytochrome P450 that bioactivates the selective pneumotoxins 3-methylindole and naphthalene.Arch Biochem Biophys. 1998 Jan 15;349(2):329-40. doi: 10.1006/abbi.1997.0479. Arch Biochem Biophys. 1998. PMID: 9448722
-
Selective dehydrogenation/oxygenation of 3-methylindole by cytochrome p450 enzymes.Drug Metab Dispos. 2001 Jul;29(7):950-3. Drug Metab Dispos. 2001. PMID: 11408359
-
Mechanism-based inactivation of lung-selective cytochrome P450 CYP2F enzymes.Drug Metab Dispos. 2008 Jan;36(1):155-62. doi: 10.1124/dmd.107.017897. Epub 2007 Oct 25. Drug Metab Dispos. 2008. PMID: 17962375
-
The metabolic basis of 3-methylindole-induced pneumotoxicity.Pharmacol Ther. 1990;46(1):105-18. doi: 10.1016/0163-7258(90)90038-4. Pharmacol Ther. 1990. PMID: 2181487 Review.
-
Porcine cytochrome 2A19 and 2E1.Basic Clin Pharmacol Toxicol. 2019 Jan;124(1):32-39. doi: 10.1111/bcpt.13121. Epub 2018 Oct 4. Basic Clin Pharmacol Toxicol. 2019. PMID: 30171805 Free PMC article. Review.
Cited by
-
Potent mutagenicity of 3-methylindole requires pulmonary cytochrome P450-mediated bioactivation: a comparison to the prototype cigarette smoke mutagens B(a)P and NNK.Chem Res Toxicol. 2010 Nov 15;23(11):1682-90. doi: 10.1021/tx100147z. Epub 2010 Aug 26. Chem Res Toxicol. 2010. PMID: 20795680 Free PMC article.
-
Intramolecular gas-phase reactions of synthetic nonheme oxoiron(IV) ions: proximity and spin-state reactivity rules.Chemistry. 2012 Sep 10;18(37):11747-60. doi: 10.1002/chem.201200105. Epub 2012 Jul 26. Chemistry. 2012. PMID: 22837063 Free PMC article.
-
Improved cytochrome P450 3A4 molecular models accurately predict the Phe215 requirement for raloxifene dehydrogenation selectivity.Biochemistry. 2010 Oct 19;49(41):9011-9. doi: 10.1021/bi101139q. Biochemistry. 2010. PMID: 20812728 Free PMC article.
-
Shape-selective interception by hydrocarbons of the O2-derived oxidant of a biomimetic nonheme iron complex.Angew Chem Int Ed Engl. 2009;48(10):1780-3. doi: 10.1002/anie.200805342. Angew Chem Int Ed Engl. 2009. PMID: 19180613 Free PMC article.
-
Directed evolution reveals requisite sequence elements in the functional expression of P450 2F1 in Escherichia coli.Chem Res Toxicol. 2012 Sep 17;25(9):1964-74. doi: 10.1021/tx300281g. Epub 2012 Aug 29. Chem Res Toxicol. 2012. PMID: 22901340 Free PMC article.
References
-
- Guengerich FP. Uncommon P450-catalyzed reactions. Curr Drug Metab. 2001;2:93–115. - PubMed
-
- Forkert PL, RP, Reid KI. Involvement of CYP2E1 and carboxylesterase enzymes in vinyl carbamate metabolism in human lung microsomes. Drug Metab Dispos. 2001;29:258–263. - PubMed
-
- Forkert PG, Lee RP. Metabolism of ethyl carbamate by pulmonary cytochrome P450 and carboxylesterase isozymes: involvement of CYP2E1 and hydrolase A. Toxicol Appl Pharmacol. 1997;146:245–254. - PubMed
-
- Raucy JL, Lasker JM, Lieber CS, Black M. Acetaminophen activation by human liver cytochromes P450IIE1 and P450IA2. Arch Biochem Biophys. 1989;271:270–283. - PubMed
-
- Rettie AE, Rettenmeier AW, Howald WN, Baillie TA. Cytochrome P-450--catalyzed formation of delta 4-VPA, a toxic metabolite of valproic acid. Science. 1987;235:890–893. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

