Structural, functional, and mutational analysis of the NblA protein provides insight into possible modes of interaction with the phycobilisome
- PMID: 18718907
- PMCID: PMC2662085
- DOI: 10.1074/jbc.M804241200
Structural, functional, and mutational analysis of the NblA protein provides insight into possible modes of interaction with the phycobilisome
Abstract
The enormous macromolecular phycobilisome antenna complex (>4 MDa) in cyanobacteria and red algae undergoes controlled degradation during certain forms of nutrient starvation. The NblA protein (approximately 6 kDa) has been identified as an essential component in this process. We have used structural, biochemical, and genetic methods to obtain molecular details on the mode of action of the NblA protein. We have determined the three-dimensional structure of the NblA protein from both the thermophilic cyanobacterium Thermosynechococcus vulcanus and the mesophilic cyanobacterium Synechococcus elongatus sp. PCC 7942. The NblA monomer has a helix-loop-helix motif which dimerizes into an open, four-helical bundle, identical to the previously determined NblA structure from Anabaena. Previous studies indicated that mutations to NblA residues near the C terminus impaired its binding to phycobilisome proteins in vitro, whereas the only mutation known to affect NblA function in vivo is located near the protein N terminus. We performed random mutagenesis of the S. elongatus nblA gene which enabled the identification of four additional amino acids crucial for NblA function in vivo. This data shows that essential amino acids are not confined to the protein termini. We also show that expression of the Anabaena nblA gene complements phycobilisome degradation in an S. elongatus NblA-null mutant despite the low homology between NblAs of these cyanobacteria. We propose that the NblA interacts with the phycobilisome via "structural mimicry" due to similarity in structural motifs found in all phycobiliproteins. This suggestion leads to a new model for the mode of NblA action which involves the entire NblA protein.
Figures

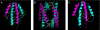



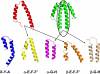

Similar articles
-
Crystal structure of NblA from Anabaena sp. PCC 7120, a small protein playing a key role in phycobilisome degradation.J Biol Chem. 2006 Feb 24;281(8):5216-23. doi: 10.1074/jbc.M507243200. Epub 2005 Dec 15. J Biol Chem. 2006. PMID: 16356935
-
A small polypeptide triggers complete degradation of light-harvesting phycobiliproteins in nutrient-deprived cyanobacteria.EMBO J. 1994 Mar 1;13(5):1039-47. doi: 10.1002/j.1460-2075.1994.tb06352.x. EMBO J. 1994. PMID: 8131738 Free PMC article.
-
NblA is essential for phycobilisome degradation in Anabaena sp. strain PCC 7120 but not for development of functional heterocysts.Microbiology (Reading). 2004 Aug;150(Pt 8):2739-2749. doi: 10.1099/mic.0.27153-0. Microbiology (Reading). 2004. PMID: 15289570
-
The phycobilisome, a light-harvesting complex responsive to environmental conditions.Microbiol Rev. 1993 Sep;57(3):725-49. doi: 10.1128/mr.57.3.725-749.1993. Microbiol Rev. 1993. PMID: 8246846 Free PMC article. Review.
-
Current Understanding of the Structure and Function of Pentapeptide Repeat Proteins.Biomolecules. 2021 Apr 26;11(5):638. doi: 10.3390/biom11050638. Biomolecules. 2021. PMID: 33925937 Free PMC article. Review.
Cited by
-
Characterization of the activities of the CpeY, CpeZ, and CpeS bilin lyases in phycoerythrin biosynthesis in Fremyella diplosiphon strain UTEX 481.J Biol Chem. 2011 Oct 14;286(41):35509-35521. doi: 10.1074/jbc.M111.284281. Epub 2011 Aug 24. J Biol Chem. 2011. PMID: 21865169 Free PMC article.
-
Structure, function, and substrates of Clp AAA+ protease systems in cyanobacteria, plastids, and apicoplasts: A comparative analysis.J Biol Chem. 2021 Jan-Jun;296:100338. doi: 10.1016/j.jbc.2021.100338. Epub 2021 Jan 23. J Biol Chem. 2021. PMID: 33497624 Free PMC article. Review.
-
Sulfate-driven elemental sparing is regulated at the transcriptional and posttranscriptional levels in a filamentous cyanobacterium.J Bacteriol. 2011 Mar;193(6):1449-60. doi: 10.1128/JB.00885-10. Epub 2011 Jan 14. J Bacteriol. 2011. PMID: 21239582 Free PMC article.
-
Structures and enzymatic mechanisms of phycobiliprotein lyases CpcE/F and PecE/F.Proc Natl Acad Sci U S A. 2017 Dec 12;114(50):13170-13175. doi: 10.1073/pnas.1715495114. Epub 2017 Nov 27. Proc Natl Acad Sci U S A. 2017. PMID: 29180420 Free PMC article.
-
Cyanophage infection in the bloom-forming cyanobacteria Microcystis aeruginosa in surface freshwater.Microbes Environ. 2012;27(4):350-5. doi: 10.1264/jsme2.me12037. Epub 2012 Oct 5. Microbes Environ. 2012. PMID: 23047146 Free PMC article. Review.
References
-
- Adir, N., Dines, M., Klartag, M., McGregor, A., and Melamed-Frank, M. (2006) in Microbiology Monographs: Inclusions in Prokaryotes (Shively, J. M., ed) pp. 47-77, Springer-Verlag, Berlin
-
- MacColl, R. (1998) J. Struct. Biol. 124 311-334 - PubMed
-
- Glazer, A. N. (1989) J. Biol. Chem. 264 1-4 - PubMed
-
- Adir, N. (2005) Photosynth. Res. 85 15-32 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

