Foxa2 and MafA regulate islet-specific glucose-6-phosphatase catalytic subunit-related protein gene expression
- PMID: 18753309
- PMCID: PMC2614309
- DOI: 10.1677/JME-08-0062
Foxa2 and MafA regulate islet-specific glucose-6-phosphatase catalytic subunit-related protein gene expression
Abstract
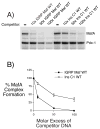
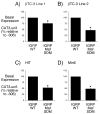
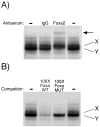
Islet-specific glucose-6-phosphatase catalytic subunit-related protein (IGRP/G6PC2) is a major autoantigen in both mouse and human type 1 diabetes. IGRP is selectively expressed in islet beta cells and polymorphisms in the IGRP gene have recently been associated with variations in fasting blood glucose levels and cardiovascular-associated mortality in humans. Chromatin immunoprecipitation (ChIP) assays have shown that the IGRP promoter binds the islet-enriched transcription factors Pax-6 and BETA2. We show here, again using ChIP assays, that the IGRP promoter also binds the islet-enriched transcription factors MafA and Foxa2. Single binding sites for these factors were identified in the proximal IGRP promoter, mutation of which resulted in decreased IGRP fusion gene expression in betaTC-3, Hamster insulinoma tumor (HIT), and Min6 cells. ChiP assays have shown that the islet-enriched transcription factor Pdx-1 also binds the IGRP promoter, but mutational analysis of four Pdx-1 binding sites in the proximal IGRP promoter revealed surprisingly little effect of Pdx-1 binding on IGRP fusion gene expression in betaTC-3 cells. In contrast, in both HIT and Min6 cells mutation of these four Pdx-1 binding sites resulted in a approximately 50% reduction in fusion gene expression. These data suggest that the same group of islet-enriched transcription factors, namely Pdx-1, Pax-6, MafA, BETA2, and Foxa2, directly or indirectly regulate expression of the two major autoantigens in type 1 diabetes.
Figures



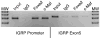



Similar articles
-
Differential regulation of islet-specific glucose-6-phosphatase catalytic subunit-related protein gene transcription by Pax-6 and Pdx-1.J Biol Chem. 2004 Aug 13;279(33):34277-89. doi: 10.1074/jbc.M404830200. Epub 2004 Jun 4. J Biol Chem. 2004. PMID: 15180990
-
Characterization of the mouse islet-specific glucose-6-phosphatase catalytic subunit-related protein gene promoter by in situ footprinting: correlation with fusion gene expression in the islet-derived betaTC-3 and hamster insulinoma tumor cell lines.Diabetes. 2001 Mar;50(3):502-14. doi: 10.2337/diabetes.50.3.502. Diabetes. 2001. PMID: 11246869
-
The proximal islet-specific glucose-6-phosphatase catalytic subunit-related protein autoantigen promoter is sufficient to initiate but not maintain transgene expression in mouse islets in vivo.Diabetes. 2004 Jul;53(7):1754-64. doi: 10.2337/diabetes.53.7.1754. Diabetes. 2004. PMID: 15220199
-
Roles and regulation of transcription factor MafA in islet beta-cells.Endocr J. 2007 Dec;54(5):659-66. doi: 10.1507/endocrj.kr-101. Epub 2007 Aug 30. Endocr J. 2007. PMID: 17785922 Review.
-
Role of PDX-1 and MafA as a potential therapeutic target for diabetes.Diabetes Res Clin Pract. 2007 Sep;77 Suppl 1:S127-37. doi: 10.1016/j.diabres.2007.01.046. Epub 2007 Apr 20. Diabetes Res Clin Pract. 2007. PMID: 17449132 Review.
Cited by
-
A single-nucleotide polymorphism in a methylatable Foxa2 binding site of the G6PC2 promoter is associated with insulin secretion in vivo and increased promoter activity in vitro.Diabetes. 2009 Feb;58(2):489-92. doi: 10.2337/db08-0587. Epub 2008 Nov 4. Diabetes. 2009. PMID: 18984742 Free PMC article.
-
Multiple promoter and enhancer differences likely contribute to augmented G6PC2 expression in human versus mouse pancreatic islet alpha cells.J Mol Endocrinol. 2024 Sep 18;73(3):e240051. doi: 10.1530/JME-24-0051. Print 2024 Oct 1. J Mol Endocrinol. 2024. PMID: 39121091
-
MafA Regulation in β-Cells: From Transcriptional to Post-Translational Mechanisms.Biomolecules. 2022 Mar 31;12(4):535. doi: 10.3390/biom12040535. Biomolecules. 2022. PMID: 35454124 Free PMC article. Review.
-
A retrospective review of the roles of multifunctional glucose-6-phosphatase in blood glucose homeostasis: Genesis of the tuning/retuning hypothesis.Life Sci. 2010 Sep 11;87(11-12):339-49. doi: 10.1016/j.lfs.2010.06.021. Epub 2010 Jul 21. Life Sci. 2010. PMID: 20603134 Free PMC article. Review.
-
An Enhancer Within Abcb11 Regulates G6pc2 in C57BL/6 Mouse Pancreatic Islets.Diabetes. 2023 Nov 1;72(11):1621-1628. doi: 10.2337/db23-0215. Diabetes. 2023. PMID: 37552875 Free PMC article.
References
-
- Ang SL, Rossant J. HNF-3 beta is essential for node and notochord formation in mouse development. Cell. 1994;78:561–574. - PubMed
-
- Aramata S, Han SI, Kataoka K. Roles and regulation of transcription factor MafA in islet beta-cells. Endocr J. 2007;54:659–666. - PubMed
-
- Arden SD, Zahn T, Steegers S, Webb S, Bergman B, O’Brien RM, Hutton JC. Molecular cloning of a pancreatic islet-specific glucose-6-phosphatase catalytic subunit-related protein. Diabetes. 1999;48:531–542. - PubMed
-
- Bischof LJ, Martin CC, Svitek CA, Stadelmaier BT, Hornbuckle LA, Goldman JK, Oeser JK, Hutton JC, O’Brien RM. Characterization of the Mouse Islet-Specific Glucose-6-Phosphatase Catalytic Subunit-Related Protein Gene Promoter by In Situ Footprinting. Correlation with Fusion Gene Expression in the Islet Derived bTC-3 and Hamster Insulinoma Tumor Cell Lines. Diabetes. 2001;50:502–514. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

