Antagonistic regulation of cell-matrix adhesion by FosB and DeltaFosB/Delta2DeltaFosB encoded by alternatively spliced forms of fosB transcripts
- PMID: 18753407
- PMCID: PMC2575163
- DOI: 10.1091/mbc.e07-08-0768
Antagonistic regulation of cell-matrix adhesion by FosB and DeltaFosB/Delta2DeltaFosB encoded by alternatively spliced forms of fosB transcripts
Abstract
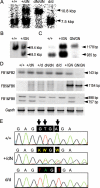
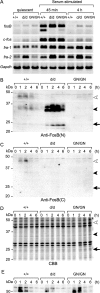
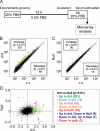
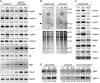
Among fos family genes encoding components of activator protein-1 complex, only the fosB gene produces two forms of mature transcripts, namely fosB and DeltafosB mRNAs, by alternative splicing of an exonic intron. The former encodes full-length FosB. The latter encodes DeltaFosB and Delta2DeltaFosB by alternative translation initiation, and both of these lack the C-terminal transactivation domain of FosB. We established two mutant mouse embryonic stem (ES) cell lines carrying homozygous fosB-null alleles and fosB(d) alleles, the latter exclusively encoding DeltaFosB/Delta2DeltaFosB. Comparison of their gene expression profiles with that of the wild type revealed that more than 200 genes were up-regulated, whereas 19 genes were down-regulated in a DeltaFosB/Delta2DeltaFosB-dependent manner. We furthermore found that mRNAs for basement membrane proteins were significantly up-regulated in fosB(d/d) but not fosB-null mutant cells, whereas genes involved in the TGF-beta1 signaling pathway were up-regulated in both mutants. Cell-matrix adhesion was remarkably augmented in fosB(d/d) ES cells and to some extent in fosB-null cells. By analyzing ES cell lines carrying homozygous fosB(FN) alleles, which exclusively encode FosB, we confirmed that FosB negatively regulates cell-matrix adhesion and the TGF-beta1 signaling pathway. We thus concluded that FosB and DeltaFosB/Delta2DeltaFosB use this pathway to antagonistically regulate cell matrix adhesion.
Figures


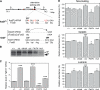

References
-
- Acquaviva C., Ferrara P., Bossis G., Brockly F., Salvat C., Jariel-Encontre I., Piechaczyk M. Degradation of cellular and viral Fos proteins. Biochimie. 2001;83:357–362. - PubMed
-
- Bertrand-Philippe M., Ruddell R. G., Arthur M. J., Thomas J., Mungalsingh N., Mann D. A. Regulation of tissue inhibitor of metalloproteinase 1 gene transcription by RUNX1 and RUNX2. J. Biol. Chem. 2004;279:24530–24539. - PubMed
-
- Boengler K., Pipp F., Fernandez B., Ziegelhoeffer T., Schaper W., Deindl E. Arteriogenesis is associated with an induction of the cardiac ankyrin repeat protein (carp) Cardiovasc. Res. 2003;59:573–581. - PubMed
-
- Botella L. M., et al. Transcriptional activation of endoglin and transforming growth factor-β signaling components by cooperative interaction between Sp1 and KLF6, their potential role in the response to vascular injury. Blood. 2002;100:4001–4010. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

