Differential loss of embryonic globin genes during the radiation of placental mammals
- PMID: 18755893
- PMCID: PMC2529089
- DOI: 10.1073/pnas.0804392105
Differential loss of embryonic globin genes during the radiation of placental mammals
Abstract
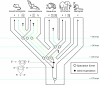
The differential gain and loss of genes from homologous gene families represents an important source of functional variation among the genomes of different species. Differences in gene content between species are primarily attributable to lineage-specific gene gains via duplication and lineage-specific losses via deletion or inactivation. Here, we use a comparative genomic approach to investigate this process of gene turnover in the beta-globin gene family of placental mammals. By analyzing genomic sequence data from representatives of each of the main superordinal clades of placental mammals, we were able to reconstruct pathways of gene family evolution during the basal radiation of this physiologically and morphologically diverse vertebrate group. Our analysis revealed that an initial expansion of the nonadult portion of the beta-globin gene cluster in the ancestor of placental mammals was followed by the differential loss and retention of ancestral gene lineages, thereby generating variation in the complement of embryonic globin genes among contemporary species. The sorting of epsilon-, gamma-, and eta-globin gene lineages among the basal clades of placental mammals has produced species differences in the functional types of hemoglobin isoforms that can be synthesized during the course of embryonic development.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Lineage-specific patterns of functional diversification in the alpha- and beta-globin gene families of tetrapod vertebrates.Mol Biol Evol. 2010 May;27(5):1126-38. doi: 10.1093/molbev/msp325. Epub 2010 Jan 4. Mol Biol Evol. 2010. PMID: 20047955 Free PMC article.
-
Platypus globin genes and flanking loci suggest a new insertional model for beta-globin evolution in birds and mammals.BMC Biol. 2008 Jul 25;6:34. doi: 10.1186/1741-7007-6-34. BMC Biol. 2008. PMID: 18657265 Free PMC article.
-
Gene Turnover and Diversification of the α- and β-Globin Gene Families in Sauropsid Vertebrates.Genome Biol Evol. 2018 Jan 1;10(1):344-358. doi: 10.1093/gbe/evy001. Genome Biol Evol. 2018. PMID: 29340581 Free PMC article.
-
Use of long sequence alignments to study the evolution and regulation of mammalian globin gene clusters.Mol Biol Evol. 1993 Jan;10(1):73-102. doi: 10.1093/oxfordjournals.molbev.a039991. Mol Biol Evol. 1993. PMID: 8383794 Review.
-
Phylogenetic diversification of the globin gene superfamily in chordates.IUBMB Life. 2011 May;63(5):313-22. doi: 10.1002/iub.482. Epub 2011 May 9. IUBMB Life. 2011. PMID: 21557448 Free PMC article. Review.
Cited by
-
CRISPR/Cas9-mediated β-globin gene knockout in rabbits recapitulates human β-thalassemia.J Biol Chem. 2021 Jan-Jun;296:100464. doi: 10.1016/j.jbc.2021.100464. Epub 2021 Feb 25. J Biol Chem. 2021. PMID: 33639162 Free PMC article.
-
Recurrent gene loss correlates with the evolution of stomach phenotypes in gnathostome history.Proc Biol Sci. 2013 Dec 4;281(1775):20132669. doi: 10.1098/rspb.2013.2669. Print 2014 Jan 22. Proc Biol Sci. 2013. PMID: 24307675 Free PMC article.
-
Gene Duplication and Evolutionary Innovations in Hemoglobin-Oxygen Transport.Physiology (Bethesda). 2016 May;31(3):223-32. doi: 10.1152/physiol.00060.2015. Physiology (Bethesda). 2016. PMID: 27053736 Free PMC article. Review.
-
Distinctive patterns of evolution of the δ-globin gene (HBD) in primates.PLoS One. 2015 Apr 8;10(4):e0123365. doi: 10.1371/journal.pone.0123365. eCollection 2015. PLoS One. 2015. PMID: 25853817 Free PMC article.
-
Copy number polymorphism in the α-globin gene cluster of European rabbit (Oryctolagus cuniculus).Heredity (Edinb). 2012 May;108(5):531-6. doi: 10.1038/hdy.2011.118. Epub 2011 Dec 7. Heredity (Edinb). 2012. PMID: 22146981 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

