Characterization of irvR, a novel regulator of the irvA-dependent pathway required for genetic competence and dextran-dependent aggregation in Streptococcus mutans
- PMID: 18757533
- PMCID: PMC2580701
- DOI: 10.1128/JB.00967-08
Characterization of irvR, a novel regulator of the irvA-dependent pathway required for genetic competence and dextran-dependent aggregation in Streptococcus mutans
Abstract
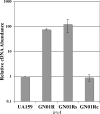
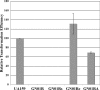
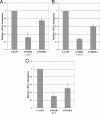
Previous studies identified irvA as a normally repressed but highly inducible transcription regulator capable of repressing mutacin I gene expression in Streptococcus mutans. In this study, we aimed to identify and characterize the regulator(s) responsible for repressing the expression of irvA. An uncharacterized open reading frame (SMU.1398) located immediately adjacent to irvA and annotated as a putative transcription repressor was identified as a likely candidate. The results of mutation studies confirmed that the expression of irvA was greatly increased in the SMU.1398 background. Mutation of SMU.1398 ("irvR") abolished genetic competence and reduced the expression of the late competence genes/operons comEA, comY, and dprA without affecting the expression of the known competence regulators comC, comED, or comX. In addition, irvR was found to be a potent negative regulator of dextran-dependent aggregation (DDAG) and gbpC expression. Each of these irvR mutant phenotypes could be rescued with a double mutation of irvA or complemented by introducing a wild-type copy of irvR on a shuttle vector. These data indicate that the repression of irvA is critically dependent upon irvR and that irvA repression is essential for the development of genetic competence and the proper control of DDAG in S. mutans.
Figures


Similar articles
-
The Streptococcus mutans IrvR repressor is a CI-like regulator that functions through autocleavage and Clp-dependent proteolysis.J Bacteriol. 2010 Mar;192(6):1586-95. doi: 10.1128/JB.01261-09. Epub 2009 Dec 28. J Bacteriol. 2010. PMID: 20038591 Free PMC article.
-
Role of the Streptococcus mutans irvA gene in GbpC-independent, dextran-dependent aggregation and biofilm formation.Appl Environ Microbiol. 2009 Nov;75(22):7037-43. doi: 10.1128/AEM.01015-09. Epub 2009 Sep 25. Appl Environ Microbiol. 2009. PMID: 19783751 Free PMC article.
-
IrvA-dependent and IrvA-independent pathways for mutacin gene regulation in Streptococcus mutans.FEMS Microbiol Lett. 2006 Aug;261(2):231-4. doi: 10.1111/j.1574-6968.2006.00351.x. FEMS Microbiol Lett. 2006. PMID: 16907725
-
SMU.940 regulates dextran-dependent aggregation and biofilm formation in Streptococcus mutans.Mol Oral Microbiol. 2018 Feb;33(1):47-58. doi: 10.1111/omi.12196. Epub 2017 Oct 11. Mol Oral Microbiol. 2018. PMID: 28845576
-
The Streptococcus mutans irvA gene encodes a trans-acting riboregulatory mRNA.Mol Cell. 2015 Jan 8;57(1):179-90. doi: 10.1016/j.molcel.2014.11.003. Mol Cell. 2015. PMID: 25574948 Free PMC article.
Cited by
-
The Streptococcus mutans IrvR repressor is a CI-like regulator that functions through autocleavage and Clp-dependent proteolysis.J Bacteriol. 2010 Mar;192(6):1586-95. doi: 10.1128/JB.01261-09. Epub 2009 Dec 28. J Bacteriol. 2010. PMID: 20038591 Free PMC article.
-
Glucan Binding Protein C of Streptococcus mutans Mediates both Sucrose-Independent and Sucrose-Dependent Adherence.Infect Immun. 2018 Jun 21;86(7):e00146-18. doi: 10.1128/IAI.00146-18. Print 2018 Jul. Infect Immun. 2018. PMID: 29685986 Free PMC article.
-
Genomewide Identification of Essential Genes and Fitness Determinants of Streptococcus mutans UA159.mSphere. 2018 Feb 7;3(1):e00031-18. doi: 10.1128/mSphere.00031-18. eCollection 2018 Jan-Feb. mSphere. 2018. PMID: 29435491 Free PMC article.
-
An Essential Role for (p)ppGpp in the Integration of Stress Tolerance, Peptide Signaling, and Competence Development in Streptococcus mutans.Front Microbiol. 2016 Jul 28;7:1162. doi: 10.3389/fmicb.2016.01162. eCollection 2016. Front Microbiol. 2016. PMID: 27516759 Free PMC article.
-
The transcription regulator BrsR serves as a network hub of natural competence protein-protein interactions in Streptococcus mutans.Proc Natl Acad Sci U S A. 2021 Sep 28;118(39):e2106048118. doi: 10.1073/pnas.2106048118. Proc Natl Acad Sci U S A. 2021. PMID: 34544866 Free PMC article.
References
-
- Ajdic, D., W. M. McShan, R. E. McLaughlin, G. Savic, J. Chang, M. B. Carson, C. Primeaux, R. Tian, S. Kenton, H. Jia, S. Lin, Y. Qian, S. Li, H. Zhu, F. Najar, H. Lai, J. White, B. A. Roe, and J. J. Ferretti. 2002. Genome sequence of Streptococcus mutans UA159, a cariogenic dental pathogen. Proc. Natl. Acad. Sci. USA 9914434-14439. - PMC - PubMed
-
- Banas, J. A. 2004. Virulence properties of Streptococcus mutans. Front. Biosci. 91267-1277. - PubMed
-
- Barsamian-Wunsch, P., J. H. Park, M. R. Watson, N. Tinanoff, and G. E. Minah. 2004. Microbiological screening for cariogenic bacteria in children 9 to 36 months of age. Pediatr. Dent. 26231-239. - PubMed
-
- Beighton, D. 2005. The complex oral microflora of high-risk individuals and groups and its role in the caries process. Community Dent. Oral Epidemiol. 33248-255. - PubMed
-
- Berge, M., M. Moscoso, M. Prudhomme, B. Martin, and J. P. Claverys. 2002. Uptake of transforming DNA in gram-positive bacteria: a view from Streptococcus pneumoniae. Mol. Microbiol. 45411-421. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

