A soluble NADH-dependent fumarate reductase in the reductive tricarboxylic acid cycle of Hydrogenobacter thermophilus TK-6
- PMID: 18757546
- PMCID: PMC2580677
- DOI: 10.1128/JB.00747-08
A soluble NADH-dependent fumarate reductase in the reductive tricarboxylic acid cycle of Hydrogenobacter thermophilus TK-6
Abstract
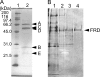
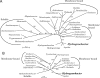
Fumarate reductase (FRD) is an enzyme that reduces fumarate to succinate. In many organisms, it is bound to the membrane and uses electron donors such as quinol. In this study, an FRD from a thermophilic chemolithoautotrophic bacterium, Hydrogenobacter thermophilus TK-6, was purified and characterized. FRD activity using NADH as an electron donor was not detected in the membrane fraction but was found in the soluble fraction. The purified enzyme was demonstrated to be a novel type of FRD, consisting of five subunits. One subunit showed high sequence identity to the catalytic subunits of known FRDs. Although the genes of typical FRDs are assembled in a cluster, the five genes encoding the H. thermophilus FRD were distant from each other in the genome. Furthermore, phylogenetic analysis showed that the H. thermophilus FRD was located in a distinct position from those of known soluble FRDs. This is the first report of a soluble NADH-dependent FRD in Bacteria and of the purification of a FRD that operates in the reductive tricarboxylic acid cycle.
Figures

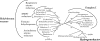
References
-
- Amino, H., H. Wang, H. Hirawake, F. Saruta, D. Mizuchi, R. Mineki, N. Shindo, K. Murayama, S. Takamiya, T. Aoki, S. Kojima, and K. Kita. 2000. Stage-specific isoforms of Ascaris suum complex II: the fumarate reductase of the parasitic adult and the succinate dehydrogenase of free-living larvae share a common iron-sulfur subunit. Mol. Biochem. Parasitol. 10663-76. - PubMed
-
- Aoshima, M. 2007. Novel enzyme reactions related to the tricarboxylic acid cycle: phylogenetic/functional implications and biotechnological applications. Appl. Microbiol. Biotechnol. 75249-255. - PubMed
-
- Beatty, J. T., and H. Gest. 1981. Generation of succinyl-coenzyme A in photosynthetic bacteria. Arch. Microbiol. 129335-340.
-
- Beh, M., G. Strauss, R. Huber, K. O. Stetter, and G. Fuchs. 1993. Enzymes of the reductive citric acid cycle in the autotrophic eubacterium Aquifex pyrophilus and in the archaebacterium Thermoproteus neutrophilus. Arch. Microbiol. 160306-311.
-
- Besteiro, S., M. Biran, N. Biteau, V. Coustou, T. Baltz, P. canioni, and F. Bringaud. 2002. Succinate secreted by Trypanosoma brucei is produced by a novel and unique glycosomal enzyme, NADH-dependent fumarate reductase. J. Biol. Chem. 27738001-38012. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

