Repeated adaptive introgression at a gene under multiallelic balancing selection
- PMID: 18769722
- PMCID: PMC2517234
- DOI: 10.1371/journal.pgen.1000168
Repeated adaptive introgression at a gene under multiallelic balancing selection
Abstract
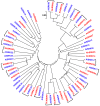
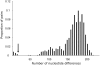
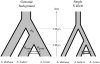
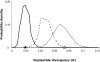
Recently diverged species typically have incomplete reproductive barriers, allowing introgression of genetic material from one species into the genomic background of the other. The role of natural selection in preventing or promoting introgression remains contentious. Because of genomic co-adaptation, some chromosomal fragments are expected to be selected against in the new background and resist introgression. In contrast, natural selection should favor introgression for alleles at genes evolving under multi-allelic balancing selection, such as the MHC in vertebrates, disease resistance, or self-incompatibility genes in plants. Here, we test the prediction that negative, frequency-dependent selection on alleles at the multi-allelic gene controlling pistil self-incompatibility specificity in two closely related species, Arabidopsis halleri and A. lyrata, caused introgression at this locus at a higher rate than the genomic background. Polymorphism at this gene is largely shared, and we have identified 18 pairs of S-alleles that are only slightly divergent between the two species. For these pairs of S-alleles, divergence at four-fold degenerate sites (K = 0.0193) is about four times lower than the genomic background (K = 0.0743). We demonstrate that this difference cannot be explained by differences in effective population size between the two types of loci. Rather, our data are most consistent with a five-fold increase of introgression rates for S-alleles as compared to the genomic background, making this study the first documented example of adaptive introgression facilitated by balancing selection. We suggest that this process plays an important role in the maintenance of high allelic diversity and divergence at the S-locus in flowering plant families. Because genes under balancing selection are expected to be among the last to stop introgressing, their comparison in closely related species provides a lower-bound estimate of the time since the species stopped forming fertile hybrids, thereby complementing the average portrait of divergence between species provided by genomic data.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
References
-
- Machado CA, Kliman RM, Markert JA, Hey J. Inferring the history of speciation from multilocus DNA sequence data: the case of Drosophila pseudoobscura and close relatives. Mol Biol Evol. 2002;19:472488. - PubMed
-
- Arnold ML. Evolution Through Genetic Exchange. (Oxford University Press, Oxford, U. K.) 2006
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Research Materials

