Insights into protein-DNA interactions through structure network analysis
- PMID: 18773096
- PMCID: PMC2518215
- DOI: 10.1371/journal.pcbi.1000170
Insights into protein-DNA interactions through structure network analysis
Abstract
Protein-DNA interactions are crucial for many cellular processes. Now with the increased availability of structures of protein-DNA complexes, gaining deeper insights into the nature of protein-DNA interactions has become possible. Earlier, investigations have characterized the interface properties by considering pairwise interactions. However, the information communicated along the interfaces is rarely a pairwise phenomenon, and we feel that a global picture can be obtained by considering a protein-DNA complex as a network of noncovalently interacting systems. Furthermore, most of the earlier investigations have been carried out from the protein point of view (protein-centric), and the present network approach aims to combine both the protein-centric and the DNA-centric points of view. Part of the study involves the development of methodology to investigate protein-DNA graphs/networks with the development of key parameters. A network representation provides a holistic view of the interacting surface and has been reported here for the first time. The second part of the study involves the analyses of these graphs in terms of clusters of interacting residues and the identification of highly connected residues (hubs) along the protein-DNA interface. A predominance of deoxyribose-amino acid clusters in beta-sheet proteins, distinction of the interface clusters in helix-turn-helix, and the zipper-type proteins would not have been possible by conventional pairwise interaction analysis. Additionally, we propose a potential classification scheme for a set of protein-DNA complexes on the basis of the protein-DNA interface clusters. This provides a general idea of how the proteins interact with the different components of DNA in different complexes. Thus, we believe that the present graph-based method provides a deeper insight into the analysis of the protein-DNA recognition mechanisms by throwing more light on the nature and the specificity of these interactions.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
 where
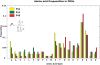
where  is the number of occurrences of the amino acid “i” in a PDG of protein–DNA complex “n” and N is the total number of structures in the dataset. The figure shows the propensities of amino acids in all the component graphs across the whole dataset.
is the number of occurrences of the amino acid “i” in a PDG of protein–DNA complex “n” and N is the total number of structures in the dataset. The figure shows the propensities of amino acids in all the component graphs across the whole dataset.




References
-
- Luscombe NM, Thornton JM. Protein–DNA interactions: amino acid conservation and the effects of mutations on binding specificity. J Mol Biol. 2002;320:991–1009. - PubMed
-
- Prabakaran P, Siebers JG, Ahmad S, Gromiha MM, Singarayan MG, et al. Classification of protein-DNA complexes based on structural descriptors. Structure. 2006;14:1355–1367. - PubMed
-
- Sathyapriya R, Brinda KV, Vishveshwara S. Correlation of the side-chain hubs with the functional residues in DNA binding protein structures. J Chem Inf Model. 2006;46:123–129. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

